|
|
AT1G48610
(Arabidopsis thaliana)
AT hook
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
Animal TF db |
| PF02178 (AT_hook) |
IPR017956 |
AT1G48610 |
T017225_2.00 |
Ensembl (2018-Dec-8) |
Link out |
| NCBI Gene Info:Predicted to encode a PR (pathogenesis-related) protein. Belongs to the lipid transfer protein (PR-14) family with the following members: At2g38540/LTP1, At2g38530/LTP2, At5g59320/LTP3, At5g59310/LTP4, At3g51600/LTP5, At3g08770/LTP6, At2g15050/LTP7, At2g18370/LTP8, At2g15325/LTP9, At5g01870/LTP10, At4g33355/LTP11, At3g51590/LTP12, At5g44265/LTP13, At5g62065/LTP14, At4g08530/LTP15. |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
| No direct experiments |
|
|
|
|
|
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
HMGA2
M09771_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$HMGA2_01
|
0.991
|
0.513
|
Hmga2
M00751_2.00 |
Mus musculus |
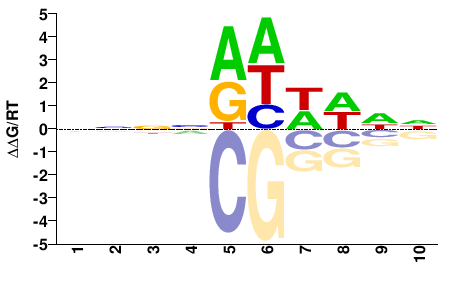
NNNNRHWWNN |
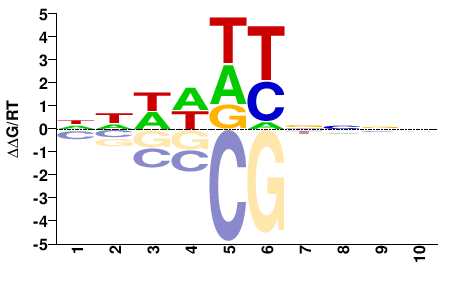
NNWWDYNNNN |
PBM
Weirauch et al.(2013)
pTH3046
|
0.991
|
0.487
|
ENSPPYG00000004736
M01685_2.00 |
Pongo abelii |
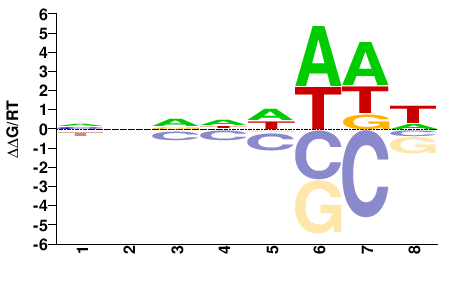
NNNNDWWW |
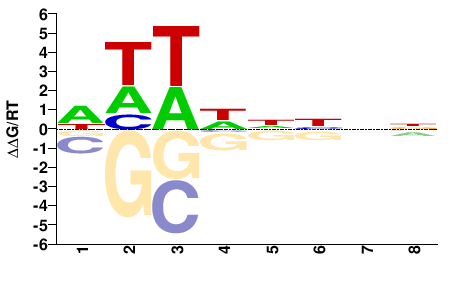
WWWHNNNN |
PBM
Weirauch et al.(2014)
pTH9279
|
0.991
|
0.487
|
HMGA1
M09769_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$HMGIY_01
|
0.991
|
0.487
|
HMGA1
M09770_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$HMGIY_Q4
|
0.991
|
0.487
|
ANIA_01690
M01675_2.00 |
Aspergillus nidulans |
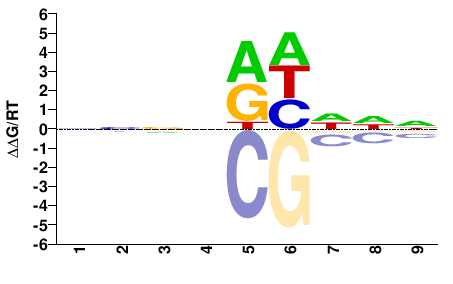
NNNNRHDDN |
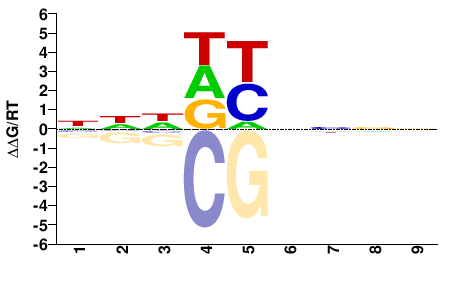
NHHDYNNNN |
PBM
Weirauch et al.(2014)
pTH8216
|
0.991
|
0.462
|
FGRRES_05233
M01104_2.00 |
Fusarium graminearum |
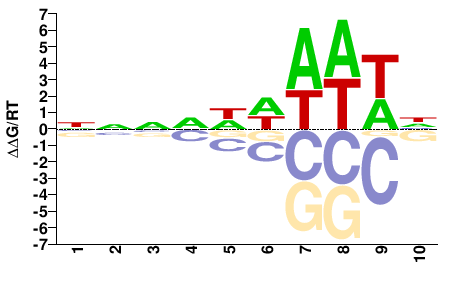
NNNDWWAWWH |
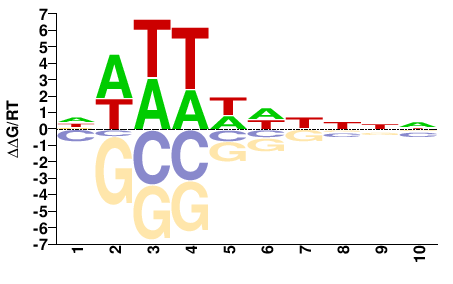
DWWTWWHNNN |
PBM
Lambert et al.(2019)
pTH11377
|
0.991
|
0.462
|
NCU05635
M01694_2.00 |
Neurospora crassa |
NNNWWWWNNN |
NNNWWWWNNN |
PBM
Weirauch et al.(2014)
pTH8863
|
0.991
|
0.436
|
HMG-I Y
M02652_2.00 |
Pisum sativum |
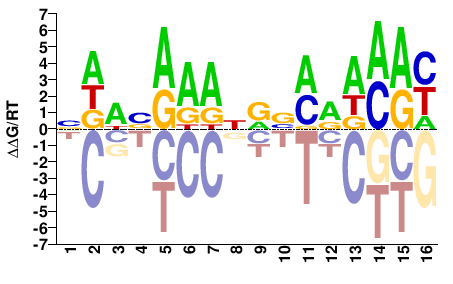

NDWVAAANRVMRWMAY |
RTKWYKBYNTTTBWHN |
SELEX
Mathelier et al.(2014)
MA0045.1
|
0.991
|
0.423
|
SUM1
M07439_2.00 |
Saccharomyces cerevisiae |

DWAAATWWN |
NWWATTTWH |
PBM, CSA and or DIP-chip
Mathelier et al.(2014)
MA0398.1
|
0.991
|
0.385
|
SUM1
M08478_2.00 |
Saccharomyces cerevisiae |

RYGWCASWAAW |

WTTWSTGWCRY |
Misc
DeBoer et al.(2011)
YDR310C_383
|
0.991
|
0.385
|
SUM1
M08479_2.00 |
Saccharomyces cerevisiae |

DWAAATWWN |
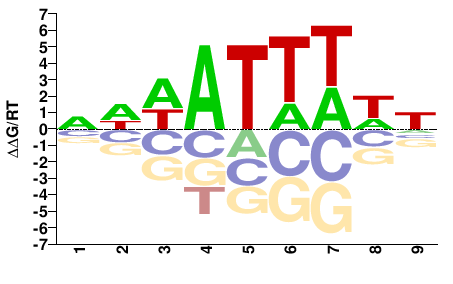
NWWATTTWH |
Misc
DeBoer et al.(2011)
YDR310C_478
|
0.991
|
0.385
|
HMGA
M01671_2.00 |
Arabidopsis thaliana |
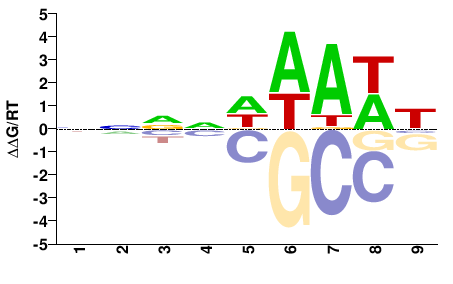
NNNNDWAWH |
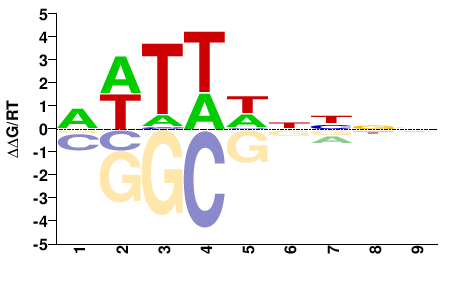
DWTWHNNNN |
PBM
Weirauch et al.(2014)
pTH7395
|
0.991
|
0.365
|
FGRRES_17512
M01105_2.00 |
Fusarium graminearum |
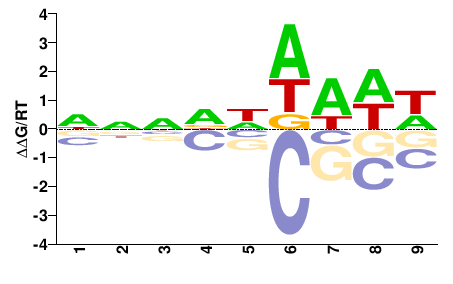
NNNDNWWWW |
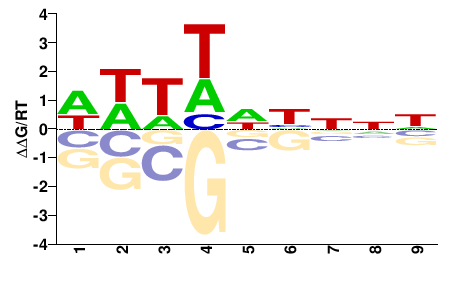
WWWWNHNNN |
PBM
Lambert et al.(2019)
pTH11412
|
0.991
|
0.359
|
athp-3
M01691_2.00 |
Caenorhabditis elegans |
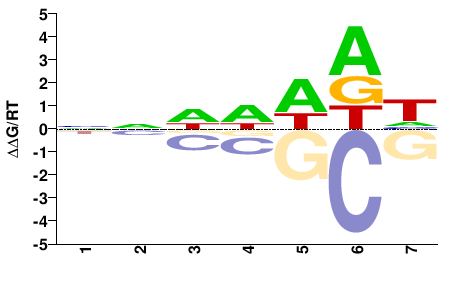
NNDDWDH |
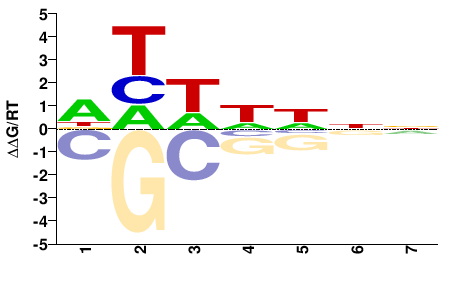
DHWHHNN |
PBM
Weirauch et al.(2014)
pTH9097
|
0.991
|
0.323
|
NCU01145
M01693_2.00 |
Neurospora crassa |
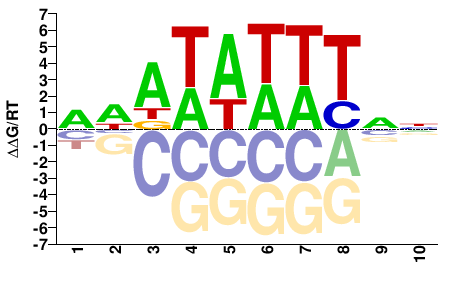
RWATAWWTNN |
NNAWWTATWY |
PBM
Weirauch et al.(2014)
pTH8916
|
0.991
|
0.323
|
TVAG_474020
M01106_2.00 |
Trichomonas vaginalis |
DNWWWAWW |
WWTWWWNH |
PBM
Lambert et al.(2019)
pTH11493
|
0.991
|
0.308
|
BRADI3G55950
M01673_2.00 |
Brachypodium distachyon |
NNNNRHDDN |
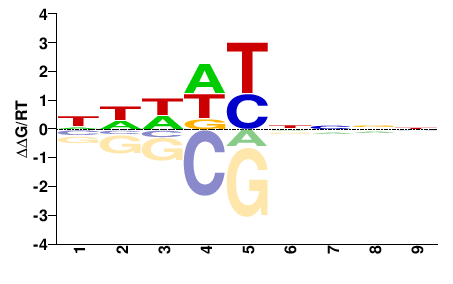
NHHDYNNNN |
PBM
Weirauch et al.(2014)
pTH9369
|
0.991
|
0.282
|
OS02G0824300
M01688_2.00 |
Oryza sativa |
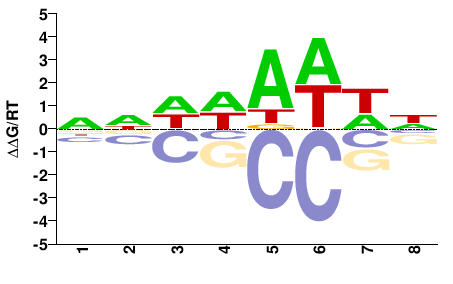
NNDWAWWN |
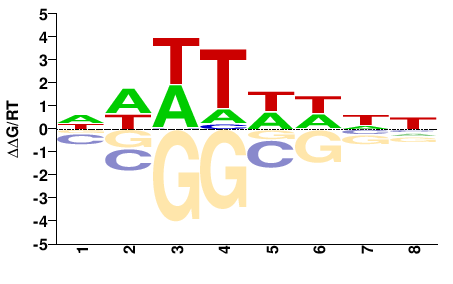
NWWTWHNN |
PBM
Weirauch et al.(2014)
pTH8406
|
0.991
|
0.282
|
Q7DLU9_PEA
M09772_2.00 |
Pisum sativum |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
P$HMGIY_01
|
0.991
|
0.282
|
AHL13
M01672_2.00 |
Arabidopsis thaliana |
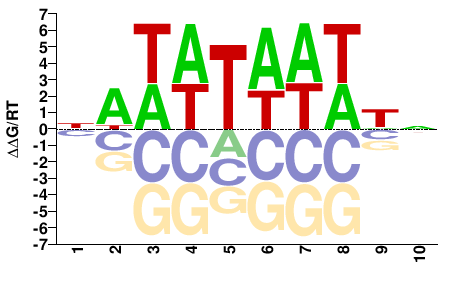
NAWWTAWWWN |
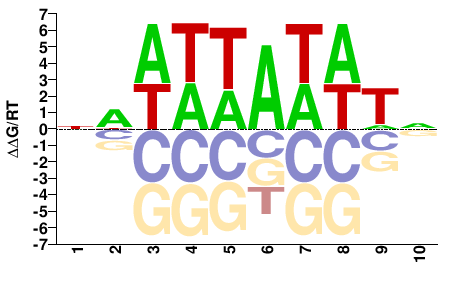
NWWWTAWWTN |
PBM
Weirauch et al.(2014)
pTH8202
|
0.991
|
0.256
|
VIT_08s0007g04020
M01689_2.00 |
Vitis vinifera |
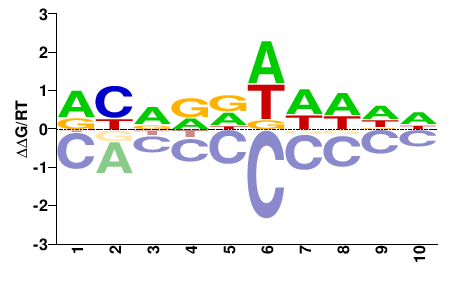
DBNDDDDDDN |
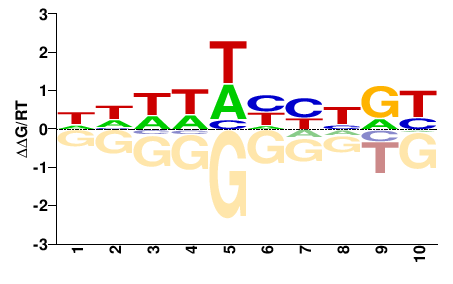
NHHHHHHNVH |
PBM
Weirauch et al.(2014)
pTH9295
|
0.991
|
0.256
|
SUM1
M00008_2.00 |
Saccharomyces cerevisiae |
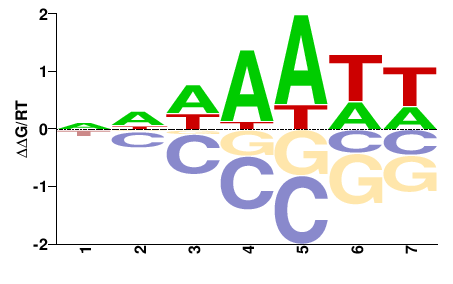
NNDWWWH |
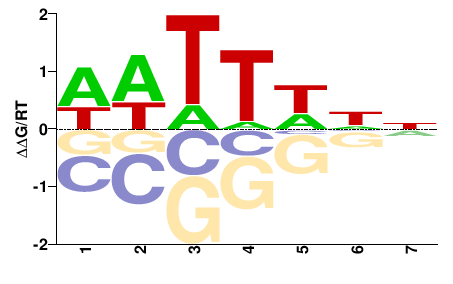
DWWWHNN |
PBM
Badis et al.(2008)
SUM1_2124
|
0.991
|
0.256
|
hmg-12
M01690_2.00 |
Caenorhabditis elegans |
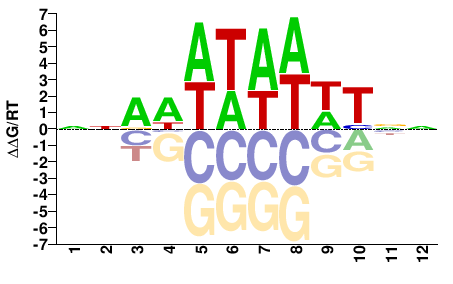
NNAWWTAWWTNN |

NNAWWTAWWTNN |
PBM
Weirauch et al.(2014)
pTH8997
|
0.991
|
0.222
|
ATEG_07399
M01676_2.00 |
Aspergillus terreus |

DHNNHNDWWW |
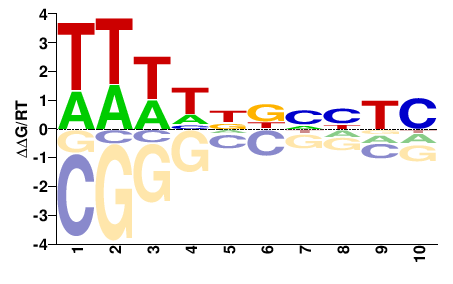
WWWHNDNNDH |
PBM
Weirauch et al.(2014)
pTH9237
|
0.991
|
0.205
|
phf21ab
M01680_2.00 |
Gasterosteus aculeatus |
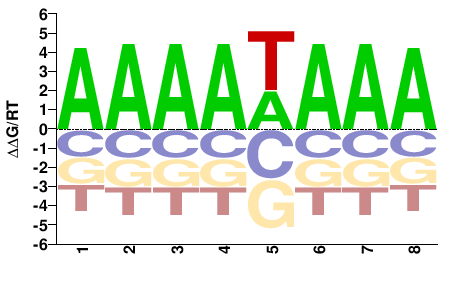
AAAAWAAA |

TTTWTTTT |
PBM
Weirauch et al.(2014)
pTH9180
|
0.991
|
0.205
|
Xrp1
M01686_2.00 |
Drosophila melanogaster |
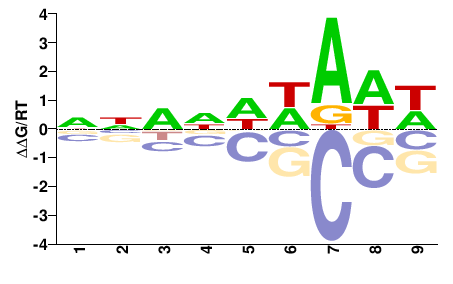
NNNNDWAWW |
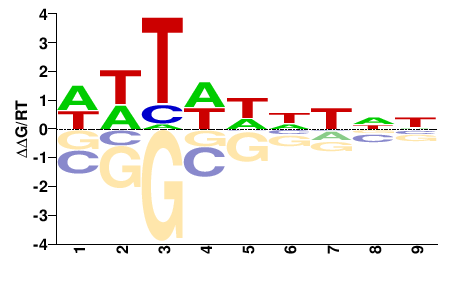
WWTWHNNNN |
PBM
Weirauch et al.(2014)
pTH9260
|
0.991
|
0.205
|
PK26376.1
M01108_2.00 |
Cannabis sativa |
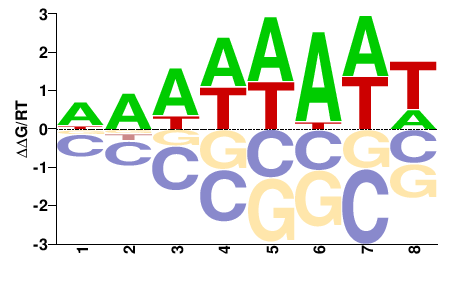
DDWWWAWW |
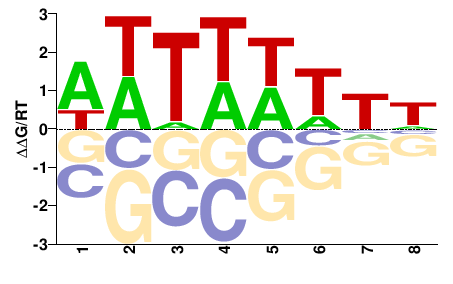

WWTWWWHH |
PBM
Lambert et al.(2019)
pTH11261
|
0.991
|
0.205
|
Xrp1
M05900_2.00 |
Drosophila melanogaster |

RTKRYGYAAT |

ATTRCRYMAY |
B1H
Zhu et al.(2011)
Xrp1_CG6272_SANGER_5_FBgn0261113
|
0.991
|
0.205
|
BRADI5G22430
M01674_2.00 |
Brachypodium distachyon |
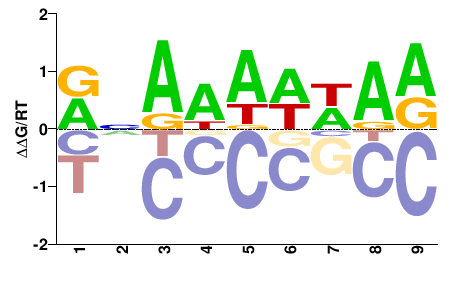
VNRDDDHDD |
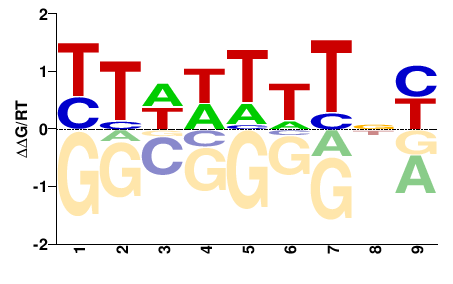

HHDHHHYNB |
PBM
Weirauch et al.(2014)
pTH9242
|
0.991
|
0.179
|
ENSACAG00000014978
M01677_2.00 |
Anolis carolinensis |
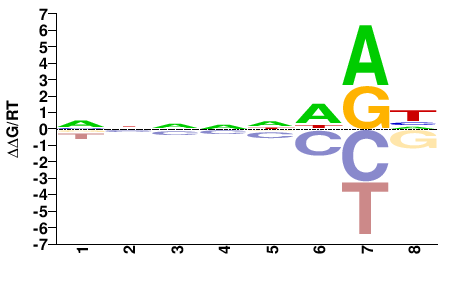
NNNNNDRH |
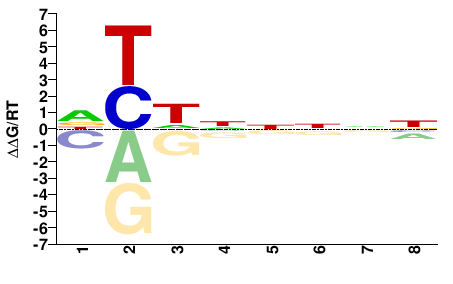
DYHNNNNN |
PBM
Weirauch et al.(2014)
pTH9222
|
0.991
|
0.179
|
phf21ab
M01678_2.00 |
Danio rerio |
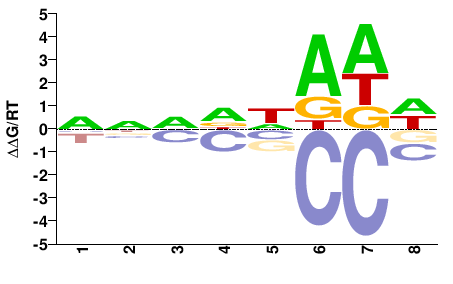
NNNDNADW |
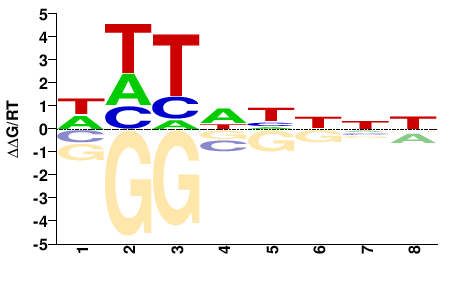
WHTNHNNN |
PBM
Weirauch et al.(2014)
pTH9254
|
0.991
|
0.179
|
kmt2cb
M01679_2.00 |
Danio rerio |

WWAWAWAT |

ATWTWTWW |
PBM
Weirauch et al.(2014)
pTH7876
|
0.991
|
0.179
|
PHF21A
M01681_2.00 |
Meleagris gallopavo |
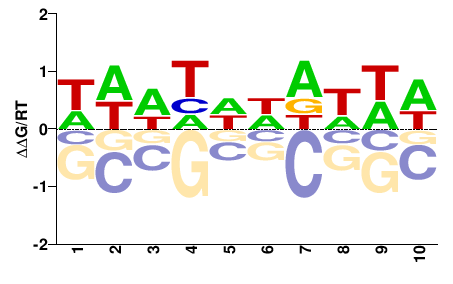

HDNHNNDNHD |
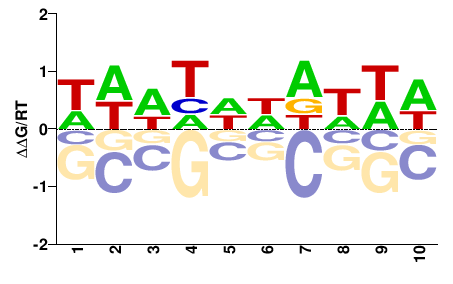

HDNHNNDNHD |
PBM
Weirauch et al.(2014)
pTH7875
|
0.991
|
0.179
|
PHF21A
M01103_2.00 |
Monodelphis domestica |
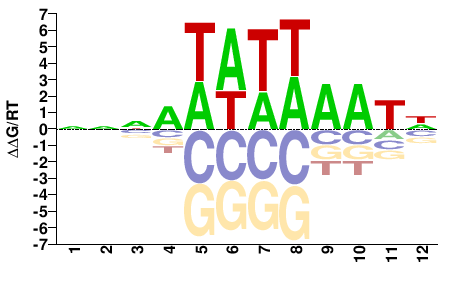
NNNAWATWAATN |
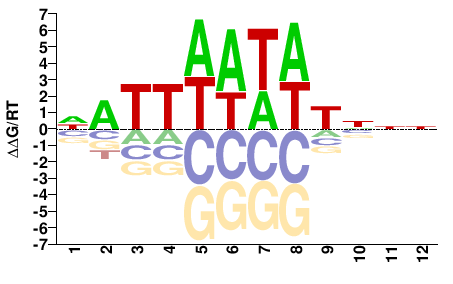

NATTWATWTNNN |
PBM
Lambert et al.(2019)
pTH9309
|
0.991
|
0.179
|
PHF21A
M01682_2.00 |
Monodelphis domestica |
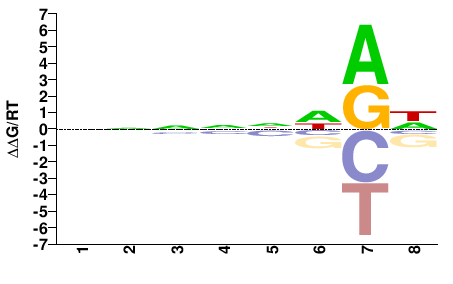
NNNNNHRH |
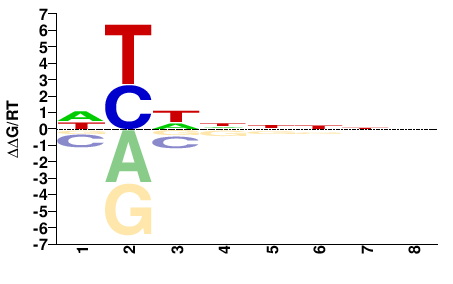
DYDNNNNN |
PBM
Weirauch et al.(2014)
pTH9335
|
0.991
|
0.179
|
Setbp1
M01683_2.00 |
Mus musculus |

NDNNNNNNHN |

NDNNNNNNHN |
PBM
Weirauch et al.(2014)
pTH1294
|
0.991
|
0.179
|
Ahctf1
M00750_2.00 |
Mus musculus |
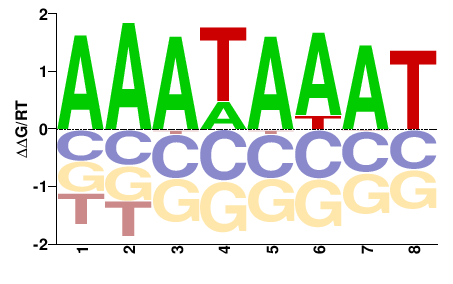

AAAWAWAW |

ATWTWTTT |
PBM
Weirauch et al.(2013)
pTH2361
|
0.991
|
0.179
|
Phf21a
M01684_2.00 |
Mus musculus |
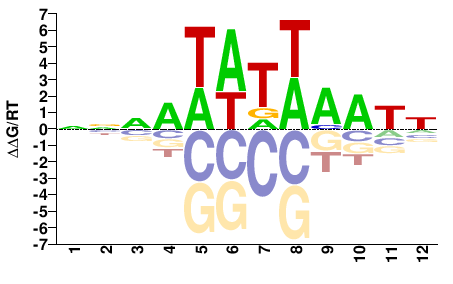
NNNATATWAATN |
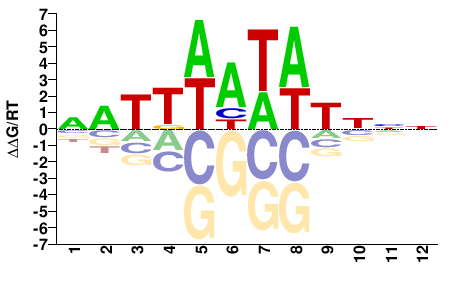
NATTWATATNNN |
PBM
Weirauch et al.(2014)
pTH8550
|
0.991
|
0.179
|
MTR_4g101990
M01687_2.00 |
Medicago truncatula |
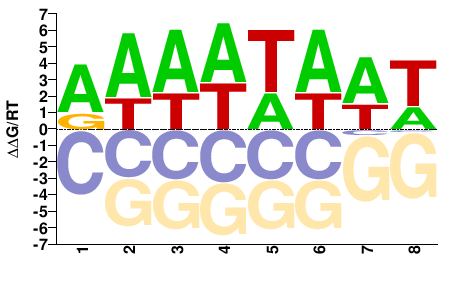
AAAWTAAT |

ATTAWTTT |
PBM
Weirauch et al.(2014)
pTH9209
|
0.991
|
0.179
|
estExt_fgeneshNG_pg.C_820014
M01692_2.00 |
Naegleria gruberi |
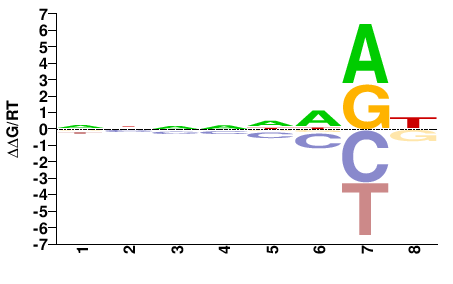
NNNNNDRH |

DYHNNNNN |
PBM
Weirauch et al.(2014)
pTH9380
|
0.991
|
0.179
|
PK00377.1
M01107_2.00 |
Cannabis sativa |
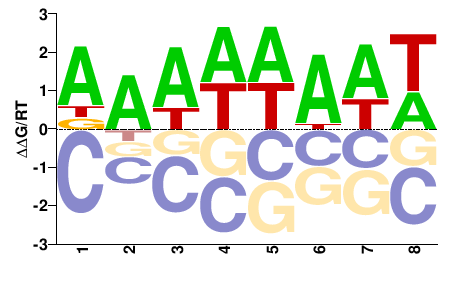
DAWWWAWW |
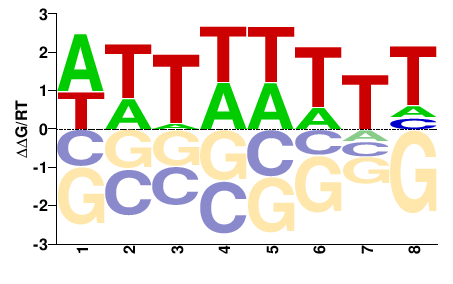

WWTWWWTH |
PBM
Lambert et al.(2019)
pTH11338
|
0.991
|
0.179
|
SUM1
M01517_2.00 |
Saccharomyces cerevisiae |
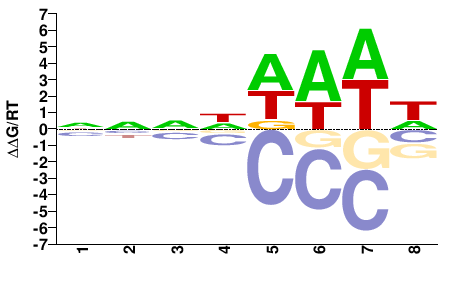
NNNDWAWW |
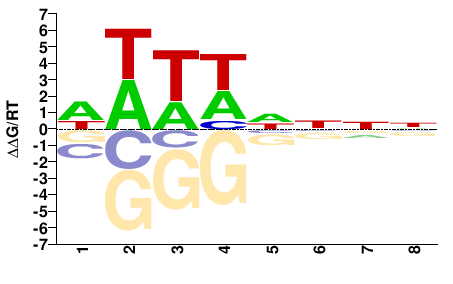
WWTWHNNN |
PBM
Zhu et al.(2009)
Sum1
|
0.991
|
0.000
|
| For this family, TFs with SR scores > 0.990 will likely have a similar motif |