|
|
HHAL001083-PA
(Halyomorpha halys)
AT hook
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
| PF02178 (AT_hook) |
IPR017956 |
HHAL001083-PA |
T018081_2.00 |
Misc (2018-Jan-19) |
| NCBI Gene Info:This gene encodes a tumor suppressor protein containing transcriptional activation, DNA binding, and oligomerization domains. The encoded protein responds to diverse cellular stresses to regulate expression of target genes, thereby inducing cell cycle arrest, apoptosis, senescence, DNA repair, or changes in metabolism. Mutations in this gene are associated with a variety of human cancers, including hereditary cancers such as Li-Fraumeni syndrome. Alternative splicing of this gene and the use of alternate promoters result in multiple transcript variants and isoforms. Additional isoforms have also been shown to result from the use of alternate translation initiation codons from identical transcript variants (PMIDs: 12032546, 20937277). [provided by RefSeq, Dec 2016] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
| No direct experiments |
|
|
|
|
|
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
BRADI3G55950
M01673_2.00 |
Brachypodium distachyon |
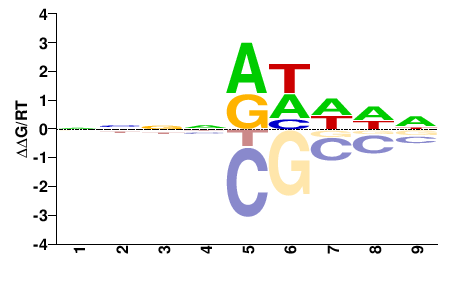
NNNNRHDDN |
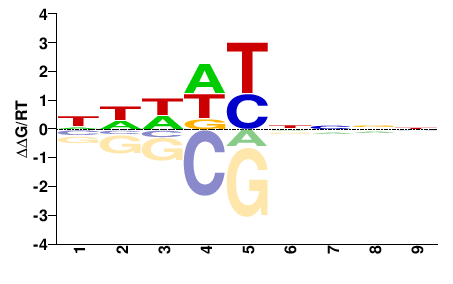
NHHDYNNNN |
PBM
Weirauch et al.(2014)
pTH9369
|
0.991
|
0.436
|
OS02G0824300
M01688_2.00 |
Oryza sativa |
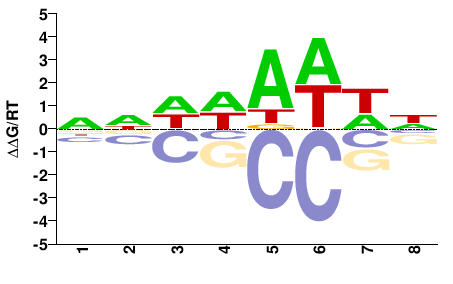
NNDWAWWN |
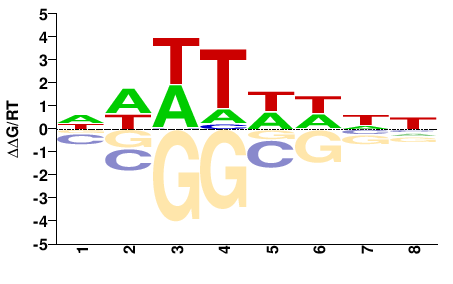
NWWTWHNN |
PBM
Weirauch et al.(2014)
pTH8406
|
0.991
|
0.436
|
NCU05635
M01694_2.00 |
Neurospora crassa |
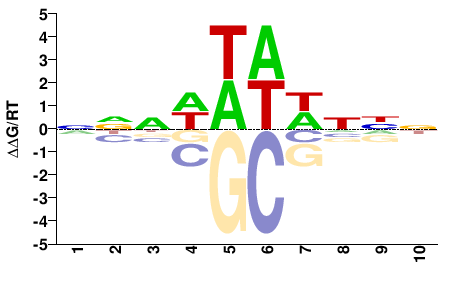
NNNWWWWNNN |
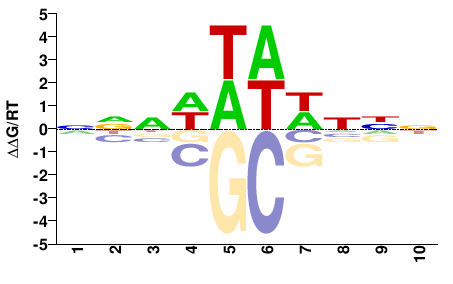
NNNWWWWNNN |
PBM
Weirauch et al.(2014)
pTH8863
|
0.991
|
0.436
|
AHL13
M01672_2.00 |
Arabidopsis thaliana |
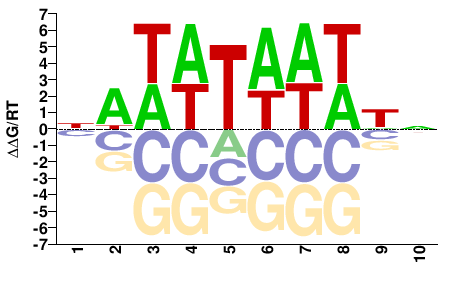
NAWWTAWWWN |
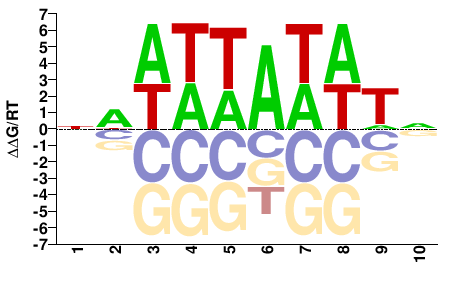
NWWWTAWWTN |
PBM
Weirauch et al.(2014)
pTH8202
|
0.991
|
0.410
|
Hmga2
M00751_2.00 |
Mus musculus |
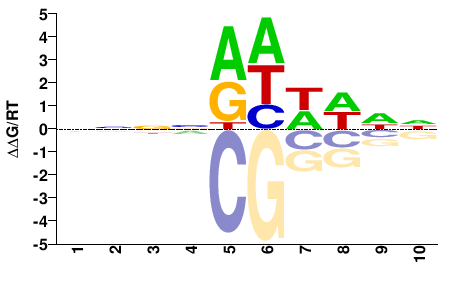
NNNNRHWWNN |
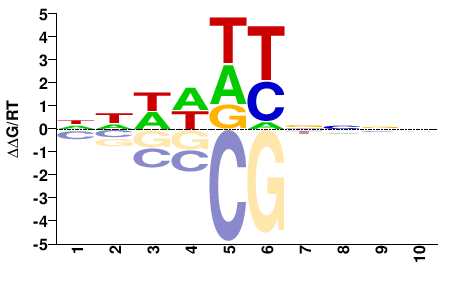
NNWWDYNNNN |
PBM
Weirauch et al.(2013)
pTH3046
|
0.991
|
0.385
|
ENSPPYG00000004736
M01685_2.00 |
Pongo abelii |
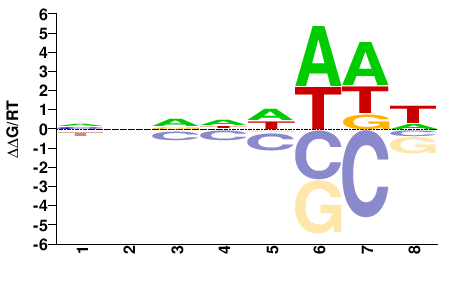
NNNNDWWW |
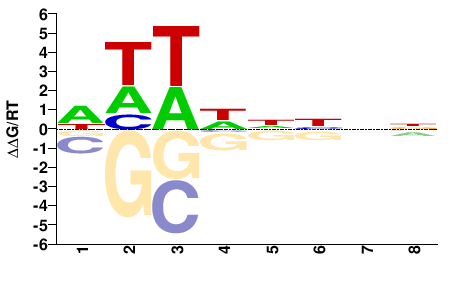
WWWHNNNN |
PBM
Weirauch et al.(2014)
pTH9279
|
0.991
|
0.385
|
FGRRES_05233
M01104_2.00 |
Fusarium graminearum |
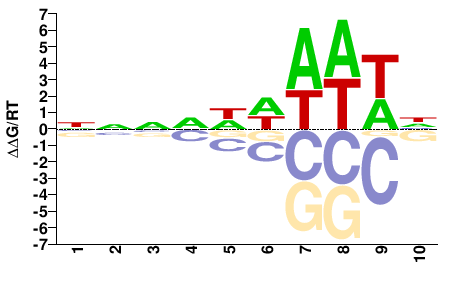
NNNDWWAWWH |
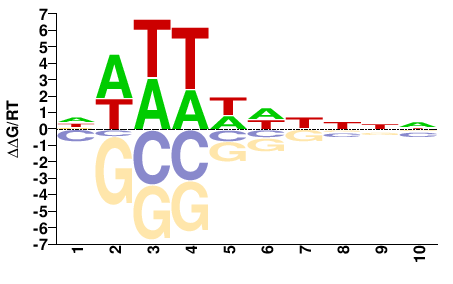
DWWTWWHNNN |
PBM
Lambert et al.(2019)
pTH11377
|
0.991
|
0.385
|
VIT_08s0007g04020
M01689_2.00 |
Vitis vinifera |
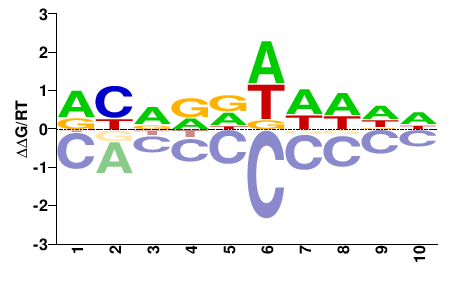
DBNDDDDDDN |
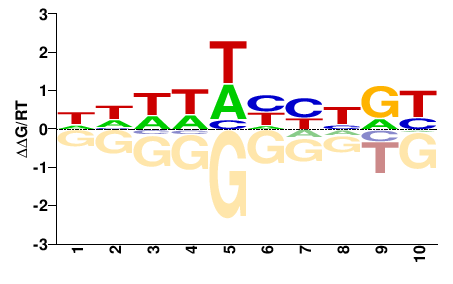
NHHHHHHNVH |
PBM
Weirauch et al.(2014)
pTH9295
|
0.991
|
0.385
|
SUM1
M07439_2.00 |
Saccharomyces cerevisiae |

DWAAATWWN |
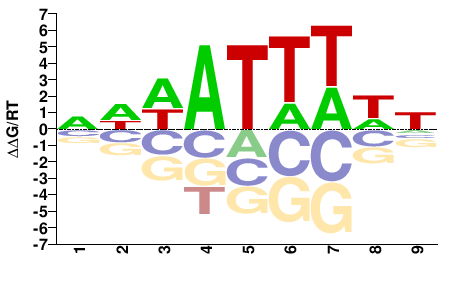
NWWATTTWH |
PBM, CSA and or DIP-chip
Mathelier et al.(2014)
MA0398.1
|
0.991
|
0.385
|
SUM1
M08478_2.00 |
Saccharomyces cerevisiae |

RYGWCASWAAW |

WTTWSTGWCRY |
Misc
DeBoer et al.(2011)
YDR310C_383
|
0.991
|
0.385
|
SUM1
M08479_2.00 |
Saccharomyces cerevisiae |

DWAAATWWN |
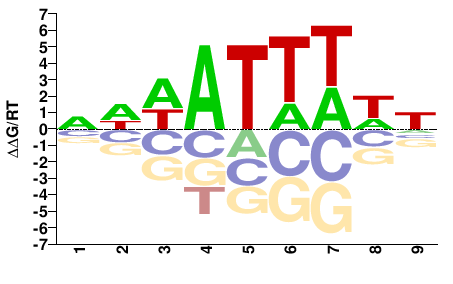
NWWATTTWH |
Misc
DeBoer et al.(2011)
YDR310C_478
|
0.991
|
0.385
|
HMGA2
M09771_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$HMGA2_01
|
0.991
|
0.385
|
HMGA1
M09769_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$HMGIY_01
|
0.991
|
0.385
|
HMGA1
M09770_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$HMGIY_Q4
|
0.991
|
0.385
|
ANIA_01690
M01675_2.00 |
Aspergillus nidulans |
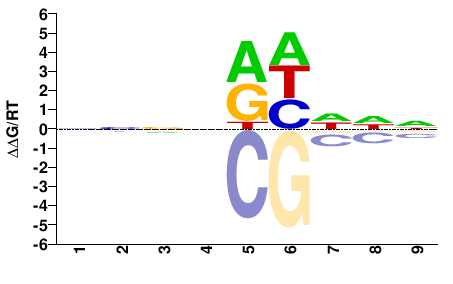
NNNNRHDDN |
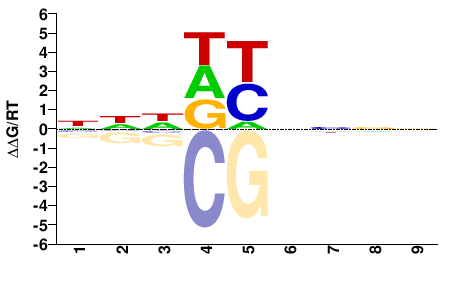
NHHDYNNNN |
PBM
Weirauch et al.(2014)
pTH8216
|
0.991
|
0.359
|
FGRRES_17512
M01105_2.00 |
Fusarium graminearum |
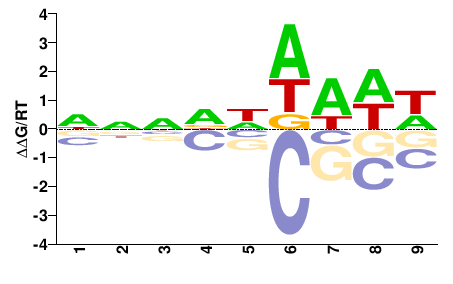
NNNDNWWWW |
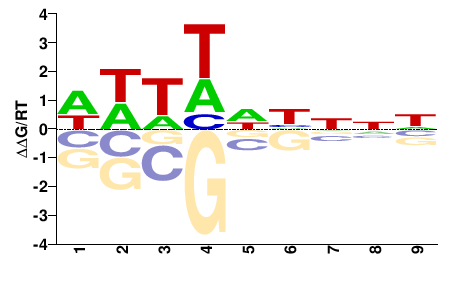
WWWWNHNNN |
PBM
Lambert et al.(2019)
pTH11412
|
0.991
|
0.308
|
TVAG_474020
M01106_2.00 |
Trichomonas vaginalis |
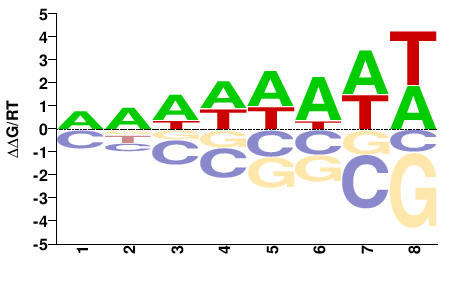
DNWWWAWW |
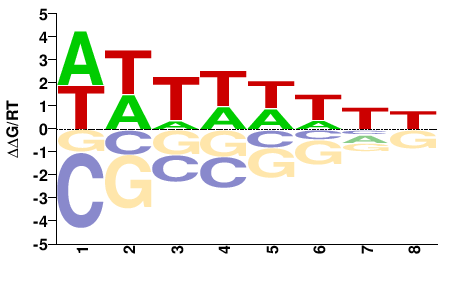
WWTWWWNH |
PBM
Lambert et al.(2019)
pTH11493
|
0.991
|
0.308
|
athp-3
M01691_2.00 |
Caenorhabditis elegans |
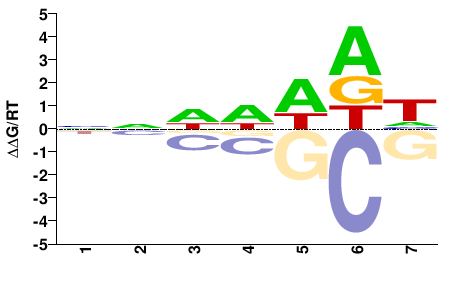
NNDDWDH |
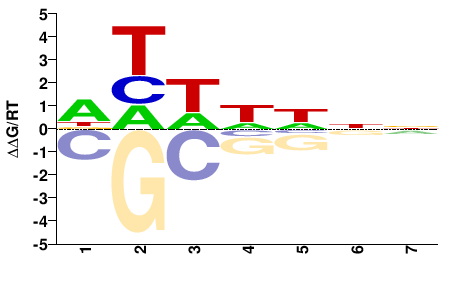
DHWHHNN |
PBM
Weirauch et al.(2014)
pTH9097
|
0.991
|
0.306
|
HMG-I Y
M02652_2.00 |
Pisum sativum |
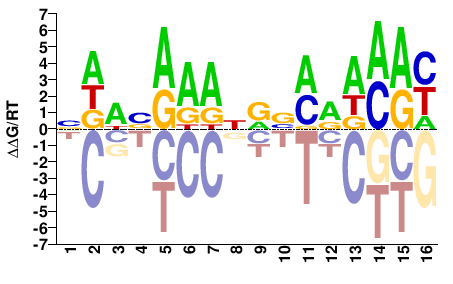

NDWVAAANRVMRWMAY |
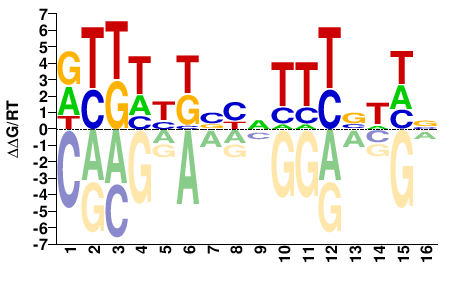
RTKWYKBYNTTTBWHN |
SELEX
Mathelier et al.(2014)
MA0045.1
|
0.991
|
0.288
|
HMGA
M01671_2.00 |
Arabidopsis thaliana |
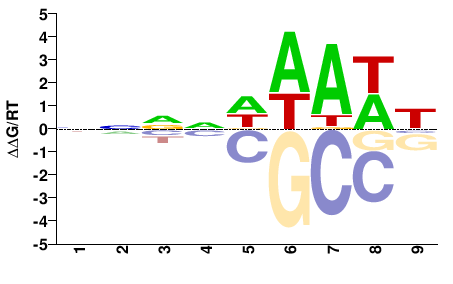
NNNNDWAWH |
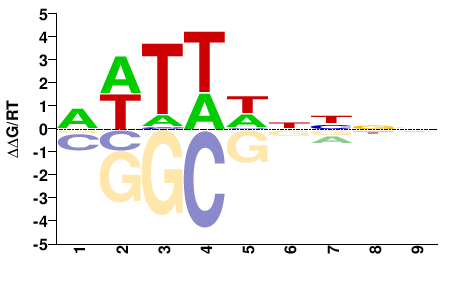
DWTWHNNNN |
PBM
Weirauch et al.(2014)
pTH7395
|
0.991
|
0.269
|
NCU01145
M01693_2.00 |
Neurospora crassa |
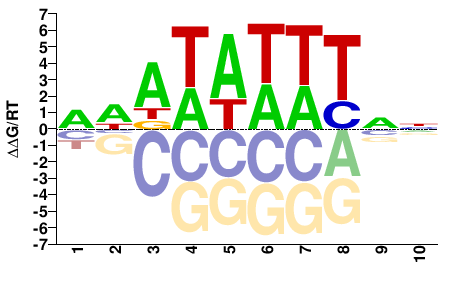
RWATAWWTNN |
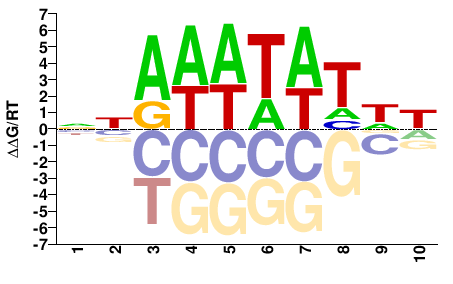
NNAWWTATWY |
PBM
Weirauch et al.(2014)
pTH8916
|
0.991
|
0.258
|
SUM1
M00008_2.00 |
Saccharomyces cerevisiae |
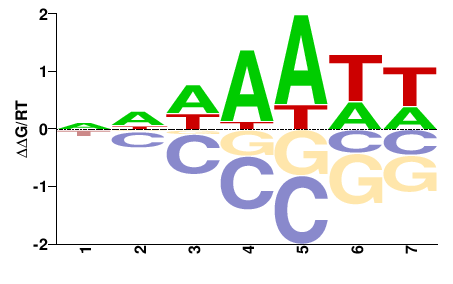
NNDWWWH |
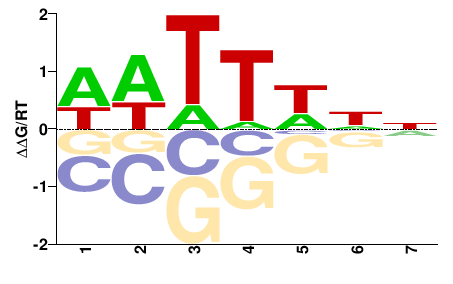
DWWWHNN |
PBM
Badis et al.(2008)
SUM1_2124
|
0.991
|
0.256
|
Q7DLU9_PEA
M09772_2.00 |
Pisum sativum |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
P$HMGIY_01
|
0.991
|
0.256
|
hmg-12
M01690_2.00 |
Caenorhabditis elegans |
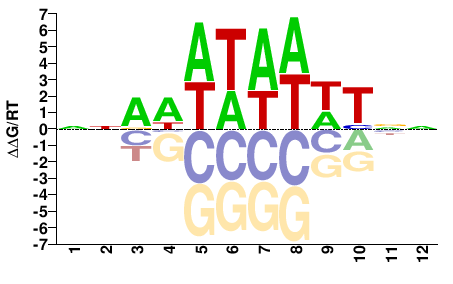
NNAWWTAWWTNN |

NNAWWTAWWTNN |
PBM
Weirauch et al.(2014)
pTH8997
|
0.991
|
0.198
|
BRADI5G22430
M01674_2.00 |
Brachypodium distachyon |
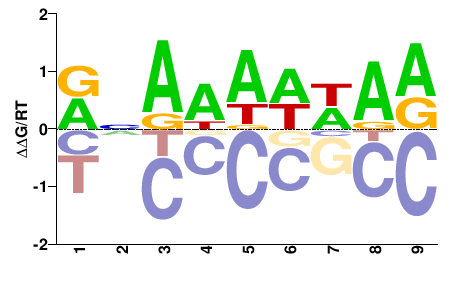
VNRDDDHDD |
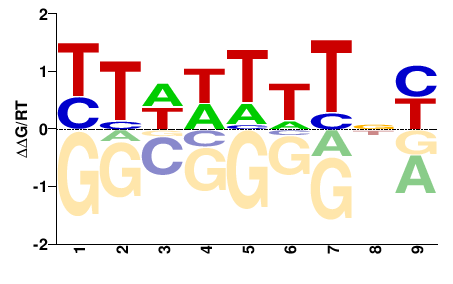

HHDHHHYNB |
PBM
Weirauch et al.(2014)
pTH9242
|
0.991
|
0.179
|
Setbp1
M01683_2.00 |
Mus musculus |

NDNNNNNNHN |

NDNNNNNNHN |
PBM
Weirauch et al.(2014)
pTH1294
|
0.991
|
0.179
|
Xrp1
M01686_2.00 |
Drosophila melanogaster |
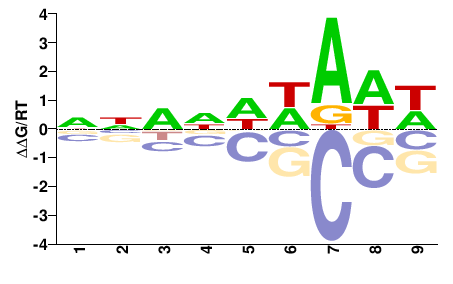
NNNNDWAWW |
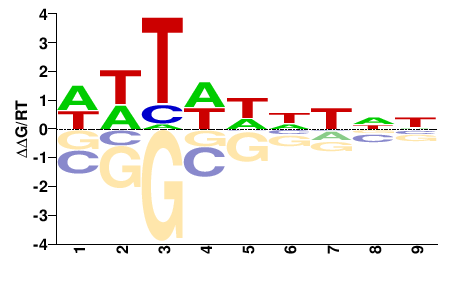
WWTWHNNNN |
PBM
Weirauch et al.(2014)
pTH9260
|
0.991
|
0.179
|
Xrp1
M05900_2.00 |
Drosophila melanogaster |

RTKRYGYAAT |

ATTRCRYMAY |
B1H
Zhu et al.(2011)
Xrp1_CG6272_SANGER_5_FBgn0261113
|
0.991
|
0.179
|
kmt2cb
M01679_2.00 |
Danio rerio |

WWAWAWAT |

ATWTWTWW |
PBM
Weirauch et al.(2014)
pTH7876
|
0.991
|
0.154
|
Ahctf1
M00750_2.00 |
Mus musculus |
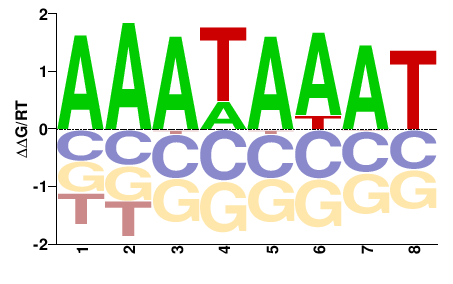

AAAWAWAW |

ATWTWTTT |
PBM
Weirauch et al.(2013)
pTH2361
|
0.991
|
0.154
|
MTR_4g101990
M01687_2.00 |
Medicago truncatula |
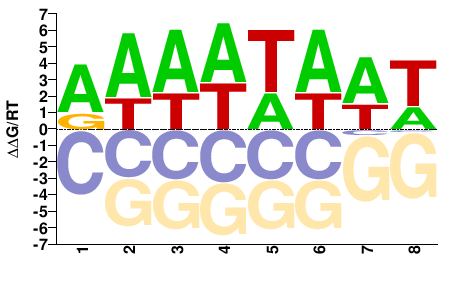
AAAWTAAT |

ATTAWTTT |
PBM
Weirauch et al.(2014)
pTH9209
|
0.991
|
0.154
|
estExt_fgeneshNG_pg.C_820014
M01692_2.00 |
Naegleria gruberi |
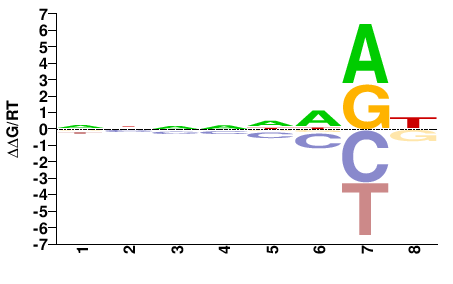
NNNNNDRH |

DYHNNNNN |
PBM
Weirauch et al.(2014)
pTH9380
|
0.991
|
0.154
|
PK00377.1
M01107_2.00 |
Cannabis sativa |
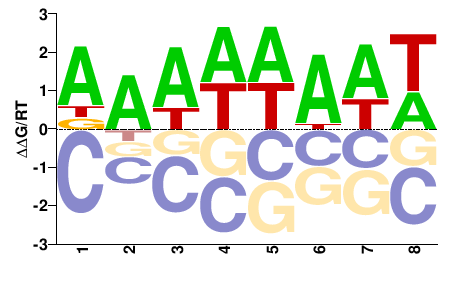
DAWWWAWW |
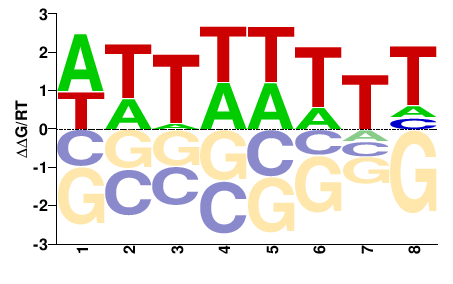

WWTWWWTH |
PBM
Lambert et al.(2019)
pTH11338
|
0.991
|
0.154
|
PK26376.1
M01108_2.00 |
Cannabis sativa |
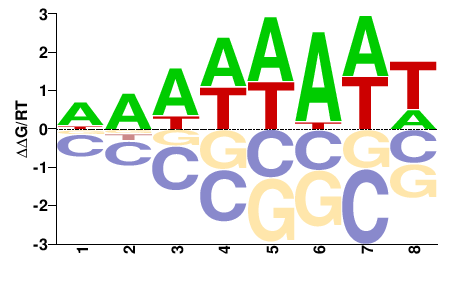
DDWWWAWW |
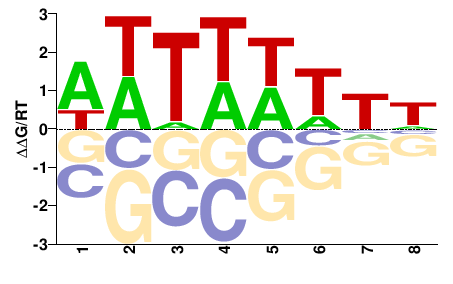

WWTWWWHH |
PBM
Lambert et al.(2019)
pTH11261
|
0.991
|
0.154
|
ATEG_07399
M01676_2.00 |
Aspergillus terreus |

DHNNHNDWWW |
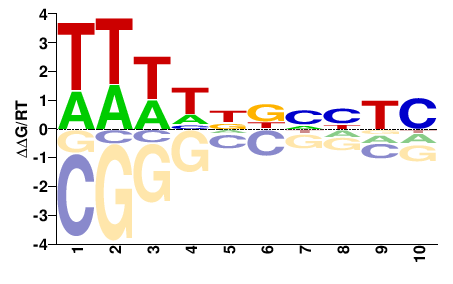
WWWHNDNNDH |
PBM
Weirauch et al.(2014)
pTH9237
|
0.991
|
0.128
|
ENSACAG00000014978
M01677_2.00 |
Anolis carolinensis |
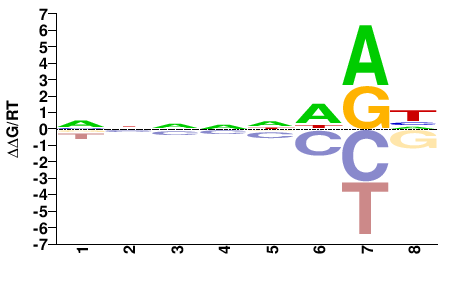
NNNNNDRH |
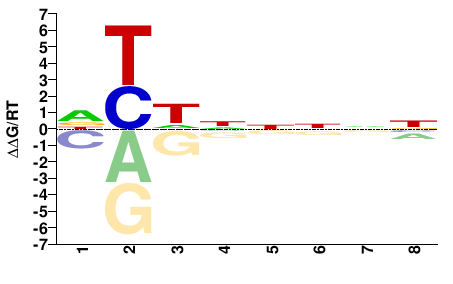
DYHNNNNN |
PBM
Weirauch et al.(2014)
pTH9222
|
0.991
|
0.128
|
phf21ab
M01678_2.00 |
Danio rerio |
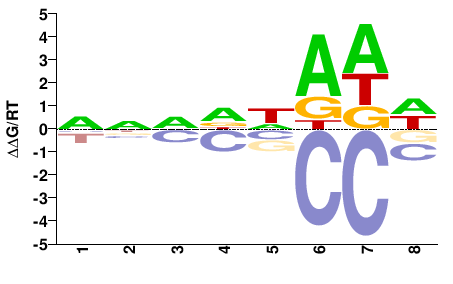
NNNDNADW |
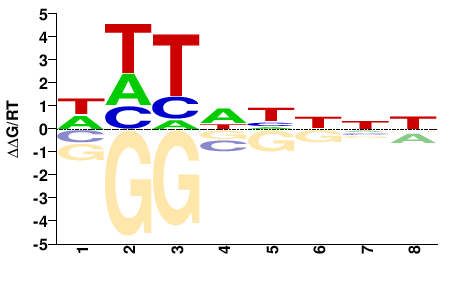
WHTNHNNN |
PBM
Weirauch et al.(2014)
pTH9254
|
0.991
|
0.128
|
phf21ab
M01680_2.00 |
Gasterosteus aculeatus |
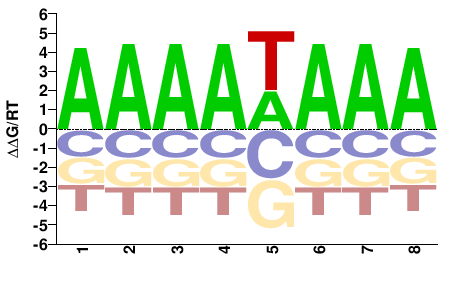
AAAAWAAA |

TTTWTTTT |
PBM
Weirauch et al.(2014)
pTH9180
|
0.991
|
0.128
|
PHF21A
M01681_2.00 |
Meleagris gallopavo |
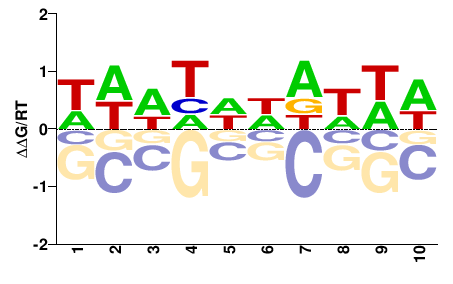

HDNHNNDNHD |
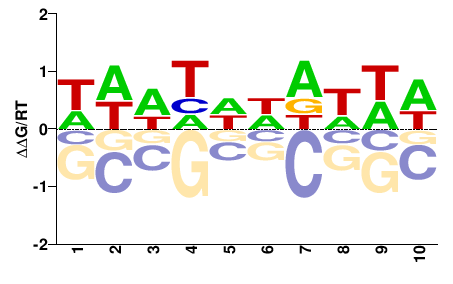

HDNHNNDNHD |
PBM
Weirauch et al.(2014)
pTH7875
|
0.991
|
0.128
|
PHF21A
M01103_2.00 |
Monodelphis domestica |
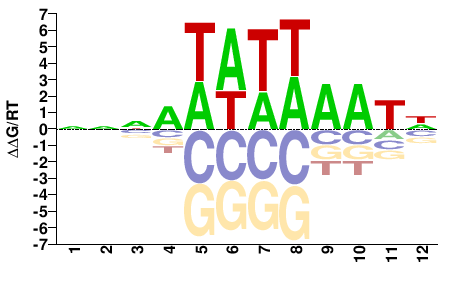
NNNAWATWAATN |
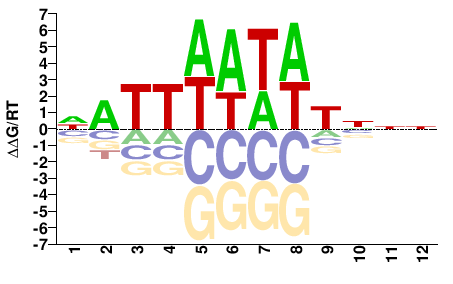

NATTWATWTNNN |
PBM
Lambert et al.(2019)
pTH9309
|
0.991
|
0.128
|
PHF21A
M01682_2.00 |
Monodelphis domestica |
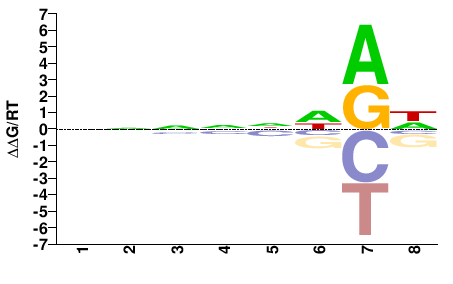
NNNNNHRH |
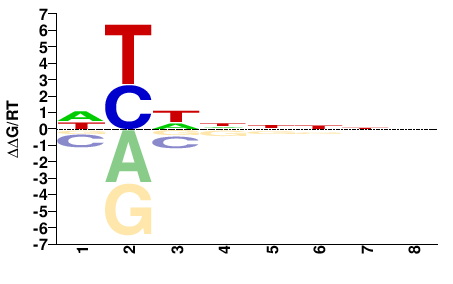
DYDNNNNN |
PBM
Weirauch et al.(2014)
pTH9335
|
0.991
|
0.128
|
Phf21a
M01684_2.00 |
Mus musculus |
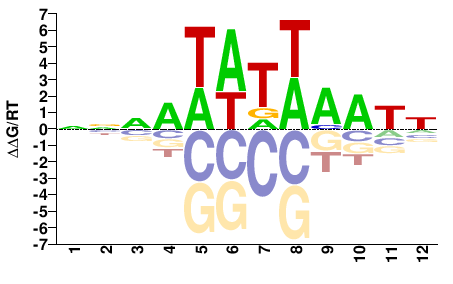
NNNATATWAATN |
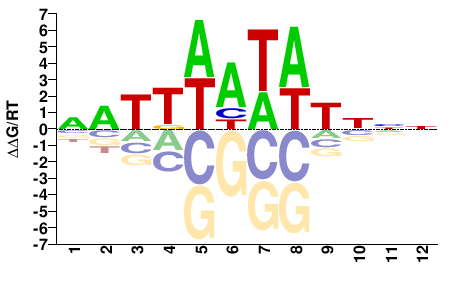
NATTWATATNNN |
PBM
Weirauch et al.(2014)
pTH8550
|
0.991
|
0.128
|
SUM1
M01517_2.00 |
Saccharomyces cerevisiae |
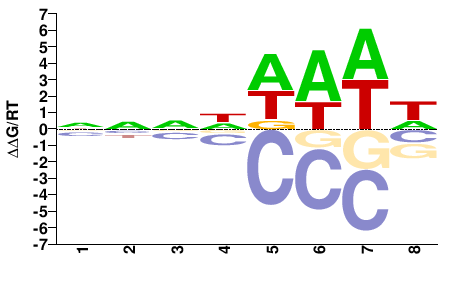
NNNDWAWW |
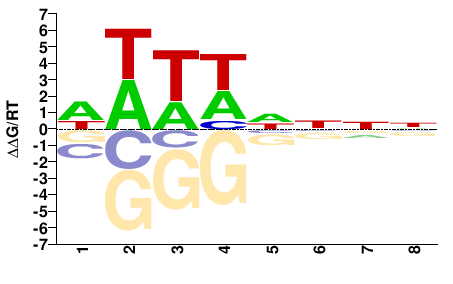
WWTWHNNN |
PBM
Zhu et al.(2009)
Sum1
|
0.991
|
0.000
|
| For this family, TFs with SR scores > 0.990 will likely have a similar motif |