|
|
Usf1
(Mus musculus)
bHLH
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
Animal TF db |
| PF00010 (HLH) |
IPR001092 |
ENSMUSG00000026641 |
T035690_2.00 |
Ensembl (2018-Dec-8) |
Link out |
| NCBI Gene Info:This protein encoded by this gene is a member of the basic-Helix-Hoop-Helix-Leucine zipper (bHLH-LZ) family and encodes a protein that can act as a transcription factor. Studies indicate that the basic region interacts with DNA at E-Box motifs, while the helix-loop-helix and leucine zipper domains are involved in dimerization with different partners. This protein is involved in a wide array of biological pathways, including cell cycle regulation, immune response, and responses to ultraviolet radiation. Mice lacking most of the coding exons of this gene often lacked both whiskers and nasal fur, and were prone to epileptic seizures, while mice lacking both this gene and another family member, Usf2, displayed embryonic lethality (PMID:9520440). Mutations in the human ortholog of this gene have been associated with Familial Combined Hyperlipidemia (FCHL) in humans. Pseudogenes of this gene are found on chromosome 11 and the X chromosome. Alternative splicing results in multiple transcript variants encoding different isoforms. [provided by RefSeq, Mar 2015] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
Usf1
M01732_2.00 |
Mus musculus |

NRYCACGTGNN |

NNCACGTGRYN |
PBM
Weirauch et al.(2014)
pTH4376 |
(Direct) |
(Direct) |
Usf1
M08756_2.00 |
Mus musculus |

NNGTCACGTGVBS |
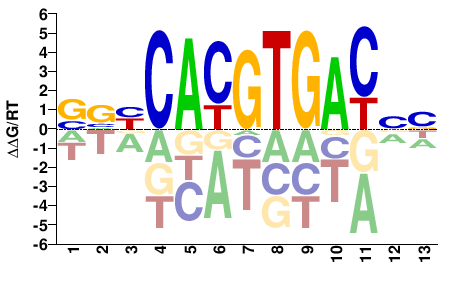

SVBCACGTGACNN |
Misc
Kulakovskiy et al.(2013)
USF1_MOUSE.H11MO.0.A |
(Direct) |
(Direct) |
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
USF1
M02792_2.00 |
Homo sapiens |

RNCACGTGRY |

RYCACGTGNY |
SELEX
Jolma et al.(2013)
USF1_1
|
0.914
|
1.000
|
USF1
M04146_2.00 |
Homo sapiens |
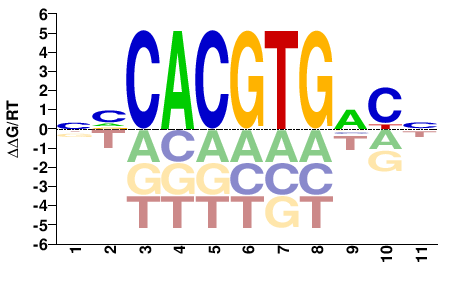
NVCACGTGVCN |

NGBCACGTGBN |
SELEX
Yin et al.(2017)
USF1_FL_HT-SELEX
|
0.914
|
1.000
|
USF1
M08056_2.00 |
Homo sapiens |

RGTCACRTGVB |

VBCAYGTGACY |
ChIP-seq
Mathelier et al.(2014)
MA0093.2
|
0.914
|
1.000
|
USF1
M07807_2.00 |
Homo sapiens |

VGTCACGTGRBSVVV |
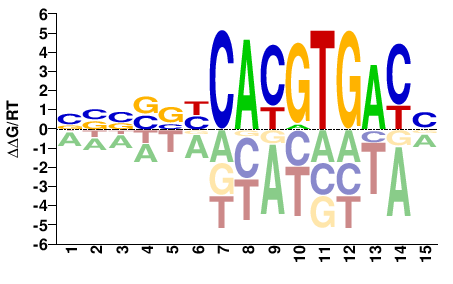
BBBSVYCACGTGACB |
ChIP-seq
Gerstein et al.(2012)
A549_USF1_HudsonAlpha
|
0.914
|
1.000
|
USF1
M07808_2.00 |
Homo sapiens |

VVGTCACGTGVBSVS |

SBSVBCACGTGACBB |
ChIP-seq
Gerstein et al.(2012)
GM12878_USF1_HudsonAlpha
|
0.914
|
1.000
|
USF1
M07809_2.00 |
Homo sapiens |

RGTCACGTGVBSVNN |

NNBSVBCACGTGACY |
ChIP-seq
Gerstein et al.(2012)
H1-hESC_USF1_HudsonAlpha
|
0.914
|
1.000
|
USF1
M07810_2.00 |
Homo sapiens |

VGTCACGTGRB |
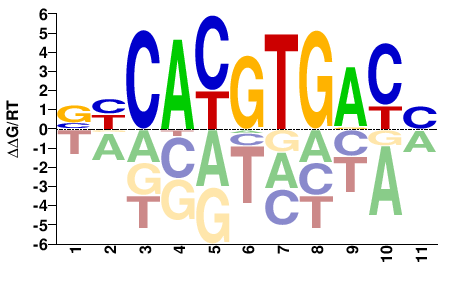

VYCACGTGACB |
ChIP-seq
Gerstein et al.(2012)
HepG2_USF1_HudsonAlpha
|
0.914
|
1.000
|
USF1
M07811_2.00 |
Homo sapiens |

NVGTCACGTGVSSV |

BSSBCACGTGACBN |
ChIP-seq
Gerstein et al.(2012)
K562_USF1_HudsonAlpha
|
0.914
|
1.000
|
USF1
M08733_2.00 |
Homo sapiens |

VGTCACGTGVBS |
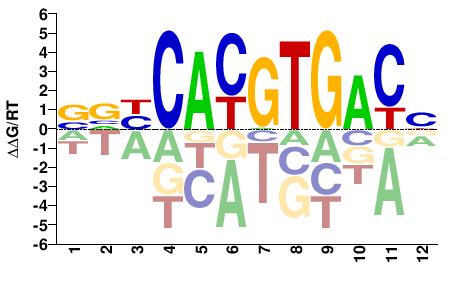

SVBCACGTGACB |
Misc
Kulakovskiy et al.(2013)
USF1_HUMAN.H11MO.0.A
|
0.914
|
1.000
|
USF1
M09460_2.00 |
Homo sapiens |

YCACGTGRBN |

NVYCACGTGR |
Misc
Heinz et al.(2010)
GM12878-Usf1_GSE32465
|
0.914
|
1.000
|
USF1
M09867_2.00 |
Homo sapiens |
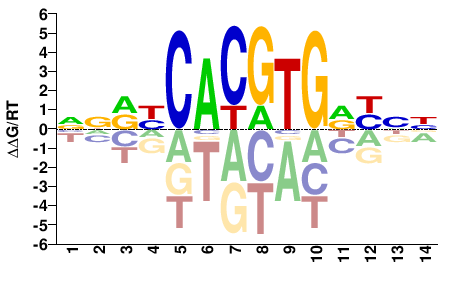
NNRYCACGTGRYNN |
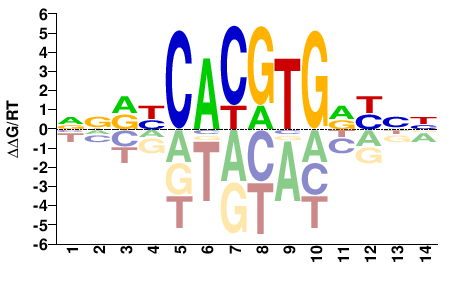
NNRYCACGTGRYNN |
Transfac
Matys et al.(2006)
V$USF_01
|
0.914
|
1.000
|
USF1
M09868_2.00 |
Homo sapiens |
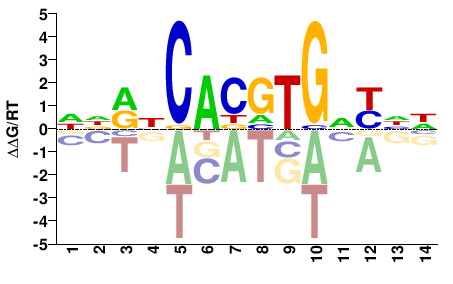
NNRNCAYRTGNYNN |
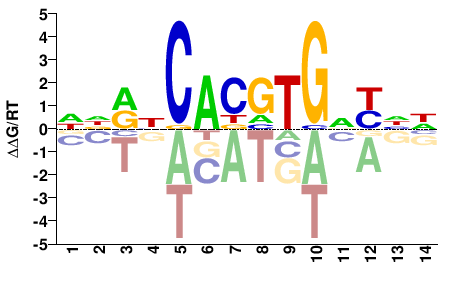
NNRNCAYRTGNYNN |
Transfac
Matys et al.(2006)
V$USF_02
|
0.914
|
1.000
|
USF1
M09869_2.00 |
Homo sapiens |

HCACGTGH |

DCACGTGD |
Transfac
Matys et al.(2006)
V$USF_C
|
0.914
|
1.000
|
USF1
M04147_2.00 |
Homo sapiens |

VHCAYGTGAYN |

NRTCACRTGDB |
SELEX
Yin et al.(2017)
USF1_FL_Methyl-HT-SELEX
|
0.914
|
1.000
|
| For this family, TFs with SR scores > 0.838 will likely have a similar motif |