|
|
ENSBTAP00000040772-D1
(Bos grunniens)
bHLH
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
| PF00010 (HLH) |
IPR001092 |
ENSBTAP00000040772-D1 |
T047156_2.00 |
GigaDB (2015-Oct-22) |
| NCBI Gene Info:This gene encodes a tumor suppressor protein containing transcriptional activation, DNA binding, and oligomerization domains. The encoded protein responds to diverse cellular stresses to regulate expression of target genes, thereby inducing cell cycle arrest, apoptosis, senescence, DNA repair, or changes in metabolism. Mutations in this gene are associated with a variety of human cancers, including hereditary cancers such as Li-Fraumeni syndrome. Alternative splicing of this gene and the use of alternate promoters result in multiple transcript variants and isoforms. Additional isoforms have also been shown to result from the use of alternate translation initiation codons from identical transcript variants (PMIDs: 12032546, 20937277). [provided by RefSeq, Dec 2016] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
| No direct experiments |
|
|
|
|
|
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
Bhlhe41
M01739_2.00 |
Mus musculus |
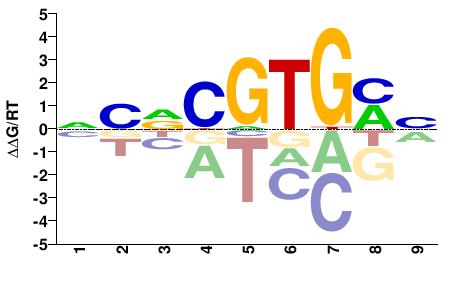
NMNCGTGMN |
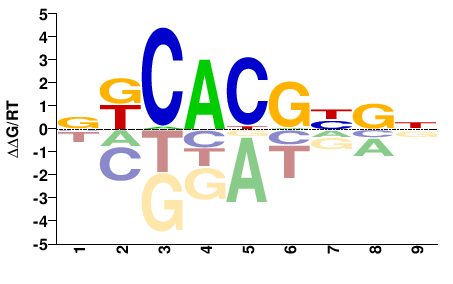

NKCACGNKN |
PBM
Weirauch et al.(2014)
pTH5060
|
0.914
|
1.000
|
BHLHE41
M02782_2.00 |
Homo sapiens |

NKCACGTGMN |

NKCACGTGMN |
SELEX
Jolma et al.(2013)
BHLHB3_1
|
0.914
|
1.000
|
BHLHE41
M02783_2.00 |
Homo sapiens |

RKCACGTGAY |

RTCACGTGMY |
SELEX
Jolma et al.(2013)
BHLHE41_1
|
0.914
|
1.000
|
BHLHE41
M04106_2.00 |
Homo sapiens |

VTCACGTGAB |

VTCACGTGAB |
SELEX
Yin et al.(2017)
BHLHE41_eDBD_HT-SELEX
|
0.914
|
1.000
|
BHLHE41
M09833_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$DEC2_Q2
|
0.914
|
1.000
|
BHLHE41
M09834_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$DEC2_Q3
|
0.914
|
1.000
|
BHLHE41
M04107_2.00 |
Homo sapiens |

RTCACGTGAY |

RTCACGTGAY |
SELEX
Yin et al.(2017)
BHLHE41_eDBD_Methyl-HT-SELEX
|
0.914
|
1.000
|
Bhlhe40
M00119_2.00 |
Mus musculus |
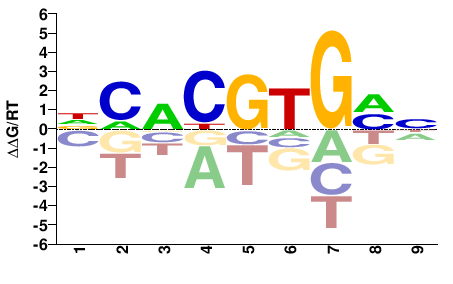
DCACGTGMN |
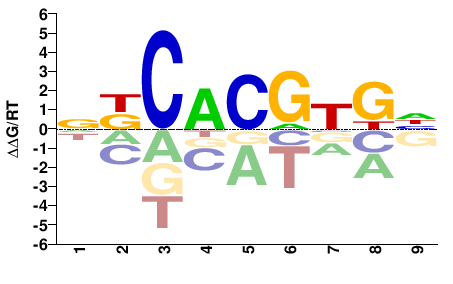

NKCACGTGH |
PBM
Badis et al.(2009)
Bhlhb2_1274
|
0.914
|
0.981
|
Bhlhe40
M01738_2.00 |
Mus musculus |
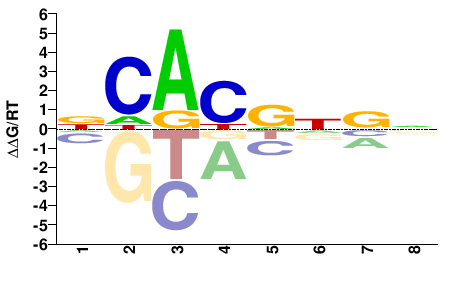
NCACRNBN |
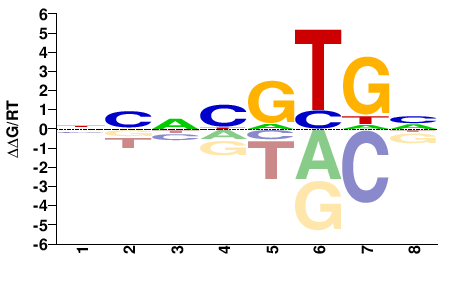
NVNYGTGN |
PBM
Weirauch et al.(2014)
pTH4330
|
0.914
|
0.981
|
BHLHE40
M02788_2.00 |
Homo sapiens |

NKCACGTGMH |

DKCACGTGMN |
SELEX
Jolma et al.(2013)
BHLHB2_1
|
0.914
|
0.981
|
Bhlhe40
M02817_2.00 |
Mus musculus |

NKCACGTGMN |

NKCACGTGMN |
SELEX
Jolma et al.(2013)
Bhlhb2_1
|
0.914
|
0.981
|
Bhlhe40
M02818_2.00 |
Mus musculus |

NKCACGTGMN |

NKCACGTGMN |
SELEX
Jolma et al.(2013)
Bhlhb2_2
|
0.914
|
0.981
|
BHLHE40
M04126_2.00 |
Homo sapiens |

VTCACGTGAB |

VTCACGTGAB |
SELEX
Yin et al.(2017)
BHLHE40_eDBD_HT-SELEX
|
0.914
|
0.981
|
BHLHE40
M07798_2.00 |
Homo sapiens |

DCACGTGMSN |
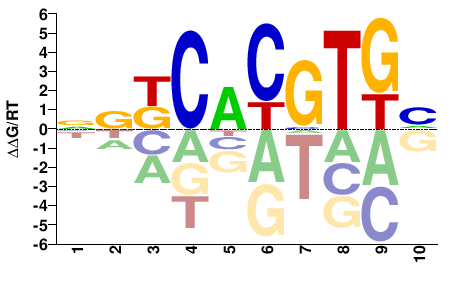

NSKCACGTGH |
ChIP-seq
Gerstein et al.(2012)
HepG2_BHLHE40_HudsonAlpha
|
0.914
|
0.981
|
BHLHE40
M08726_2.00 |
Homo sapiens |

DDCACGTGMS |

SKCACGTGHH |
Misc
Kulakovskiy et al.(2013)
BHE40_HUMAN.H11MO.0.A
|
0.914
|
0.981
|
Bhlhe40
M08761_2.00 |
Mus musculus |

DCACGTGMS |
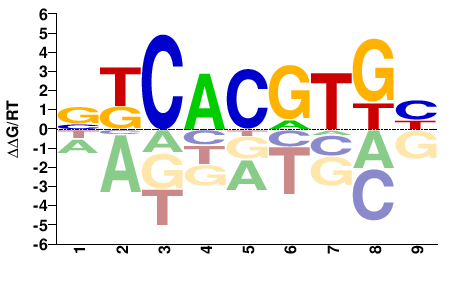

SKCACGTGH |
Misc
Kulakovskiy et al.(2013)
BHE40_MOUSE.H11MO.0.A
|
0.914
|
0.981
|
BHLHE40
M09456_2.00 |
Homo sapiens |
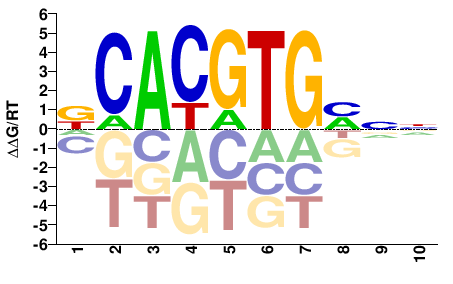
DCACGTGMNN |

NNKCACGTGH |
Misc
Heinz et al.(2010)
HepG2-BHLHE40_GSE31477
|
0.914
|
0.981
|
BHLHE40
M09842_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$BHLHE40_03
|
0.914
|
0.981
|
BHLHE40
M09843_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$DEC1_Q2
|
0.914
|
0.981
|
BHLHE40
M09844_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$DEC1_Q3
|
0.914
|
0.981
|
BHLHE40
M09845_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$STRA13_01
|
0.914
|
0.981
|
BHLHE40
M04127_2.00 |
Homo sapiens |

RTCACGTGAY |

RTCACGTGAY |
SELEX
Yin et al.(2017)
BHLHE40_eDBD_Methyl-HT-SELEX
|
0.914
|
0.981
|
| For this family, TFs with SR scores > 0.838 will likely have a similar motif |