|
|
HHAL004914-PA
(Halyomorpha halys)
Forkhead
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
| PF00250 (Forkhead) |
IPR001766 |
HHAL004914-PA |
T189529_2.00 |
Misc (2018-Jan-19) |
| NCBI Gene Info:This gene encodes a tumor suppressor protein containing transcriptional activation, DNA binding, and oligomerization domains. The encoded protein responds to diverse cellular stresses to regulate expression of target genes, thereby inducing cell cycle arrest, apoptosis, senescence, DNA repair, or changes in metabolism. Mutations in this gene are associated with a variety of human cancers, including hereditary cancers such as Li-Fraumeni syndrome. Alternative splicing of this gene and the use of alternate promoters result in multiple transcript variants and isoforms. Additional isoforms have also been shown to result from the use of alternate translation initiation codons from identical transcript variants (PMIDs: 12032546, 20937277). [provided by RefSeq, Dec 2016] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
| No direct experiments |
|
|
|
|
|
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
fkh
M03734_2.00 |
Drosophila melanogaster |

WRWGYMAATATTKRCWYW |

WRWGYMAATATTKRCWYW |
SELEX
Nitta et al.(2015)
fkh_1
|
0.855
|
0.942
|
fkh
M03735_2.00 |
Drosophila melanogaster |

HWASAATAAYAWT |
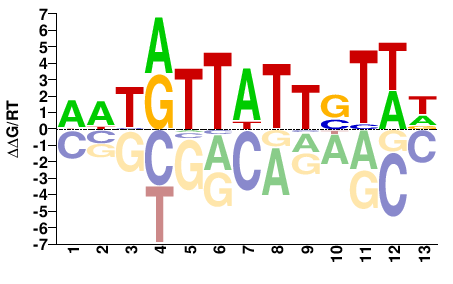
AATRTTATTSTWD |
SELEX
Nitta et al.(2015)
fkh_2
|
0.855
|
0.942
|
fkh
M06473_2.00 |
Drosophila melanogaster |

NTRDDYAAACA |

TGTTTRHHYAN |
B1H
Mathelier et al.(2014)
MA0446.1
|
0.855
|
0.942
|
fkh
M06194_2.00 |
Drosophila melanogaster |

NTRDDYAAACA |

TGTTTRHHYAN |
B1H
Zhu et al.(2011)
fkh_NAR_FBgn0000659
|
0.855
|
0.942
|
fkh
M08200_2.00 |
Drosophila melanogaster |

NTRDDYAAACA |

TGTTTRHHYAN |
ChIP-seq
Contrino et al.(2012)
Mf15
|
0.855
|
0.942
|
fkh
M09668_2.00 |
Drosophila melanogaster |

TRTTTRCAHAAG |

CTTDTGYAAAYA |
Misc
Kulakovskiy et al.(2009)
fkh
|
0.855
|
0.942
|
FOXA1
M04817_2.00 |
Homo sapiens |

NTRTKTACMYWN |

NWRKGTAMAYAN |
SELEX
Yin et al.(2017)
FOXA1_eDBD_HT-SELEX
|
0.850
|
0.907
|
FOXA1
M04819_2.00 |
Homo sapiens |

BVYTAWGTAAACAAWN |

NWTTGTTTACWTARBV |
SELEX
Yin et al.(2017)
FOXA1_FL_HT-SELEX_1
|
0.850
|
0.907
|
FOXA1
M04820_2.00 |
Homo sapiens |
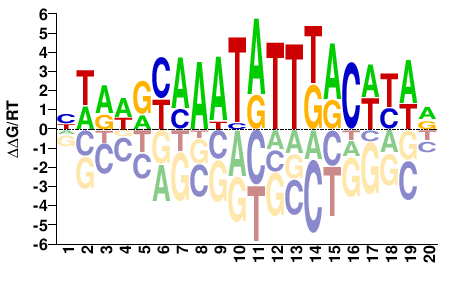
HWRWRYAAATATTKACWYWR |
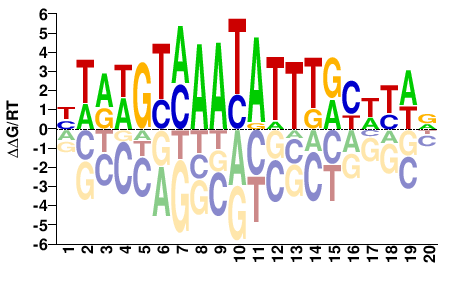

YWRWGTMAATATTTRYWYWD |
SELEX
Yin et al.(2017)
FOXA1_FL_HT-SELEX_2
|
0.850
|
0.907
|
FOXA1
M08117_2.00 |
Homo sapiens |

NNNNTRTTTACWYWD |

HWRWGTAAAYANNNN |
ChIP-seq
Mathelier et al.(2014)
MA0148.3
|
0.850
|
0.907
|
FOXA1
M07960_2.00 |
Homo sapiens |

NNYWRWGYAAACANN |

NNTGTTTRCWYWRNN |
ChIP-seq
Gerstein et al.(2012)
ECC-1_FOXA1_HudsonAlpha
|
0.850
|
0.907
|
FOXA1
M07961_2.00 |
Homo sapiens |

HWRWGTAAAYA |

TRTTTACWYWD |
ChIP-seq
Gerstein et al.(2012)
HepG2_FOXA1_HudsonAlpha
|
0.850
|
0.907
|
FOXA1
M07962_2.00 |
Homo sapiens |

NWVWGTAAACA |

TGTTTACWBWN |
ChIP-seq
Gerstein et al.(2012)
T-47D_FOXA1_HudsonAlpha
|
0.850
|
0.907
|
FOXA1
M08199_2.00 |
Homo sapiens |

BWRDGTAAACANN |

NNTGTTTACHYWV |
ChIP-seq
Contrino et al.(2012)
Mv69
|
0.850
|
0.907
|
FOXA1
M08028_2.00 |
Homo sapiens |

WRWRYAAAYA |

TRTTTRYWYW |
ChIP-seq
Chen et al.(2011)
GSE15244_FoxA1
|
0.850
|
0.907
|
FOXA1
M09090_2.00 |
Homo sapiens |

VHWRWGTAAACA |

TGTTTACWYWDB |
Misc
Kulakovskiy et al.(2013)
FOXA1_HUMAN.H11MO.0.A
|
0.850
|
0.907
|
Foxa1
M09103_2.00 |
Mus musculus |

TRTTKACWYW |
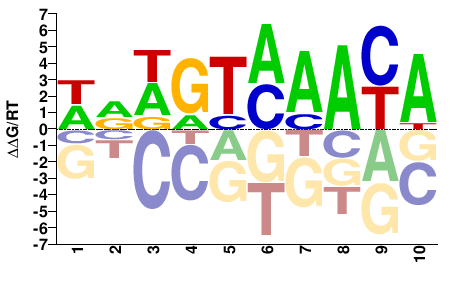
WRWGTMAAYA |
Misc
Kulakovskiy et al.(2013)
FOXA1_MOUSE.H11MO.0.A
|
0.850
|
0.907
|
FOXA1
M09544_2.00 |
Homo sapiens |
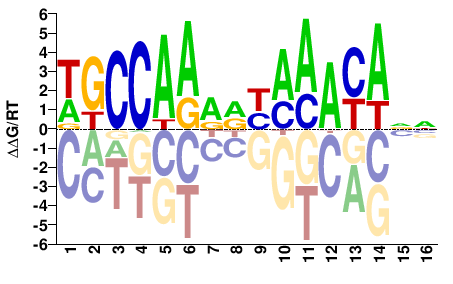
WGCCAARRYAAAYANN |
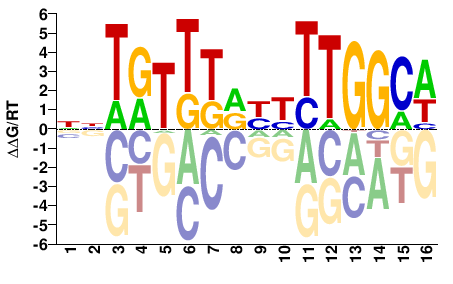
NNTRTTTRYYTTGGCW |
Misc
Heinz et al.(2010)
LNCAP-FOXA1_GSE27824_2
|
0.850
|
0.907
|
FOXA1
M09543_2.00 |
Homo sapiens |

WRWGTAAACA |

TGTTTACWYW |
Misc
Heinz et al.(2010)
LNCAP-FOXA1_GSE27824_1
|
0.850
|
0.907
|
FOXA1
M09545_2.00 |
Homo sapiens |

WRWGTAAAYA |

TRTTTACWYW |
Misc
Heinz et al.(2010)
MCF7-FOXA1_GSE26831
|
0.850
|
0.907
|
FOXA1
M10542_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$FOXA1_02
|
0.850
|
0.907
|
FOXA1
M10543_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$FOXA1_03
|
0.850
|
0.907
|
FOXA1
M10544_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$HNF3A_01
|
0.850
|
0.907
|
FOXA1
M10545_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$HNF3ALPHA_Q6
|
0.850
|
0.907
|
FOXA1
M10546_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$HNF3A_Q6
|
0.850
|
0.907
|
FOXA1
M04818_2.00 |
Homo sapiens |
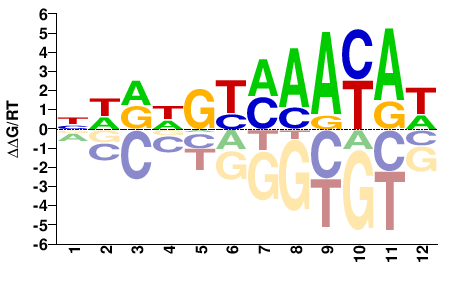
NWRDGYMAAYAW |

WTRTTKRCHYWN |
SELEX
Yin et al.(2017)
FOXA1_eDBD_Methyl-HT-SELEX
|
0.850
|
0.907
|
FOXA1
M04821_2.00 |
Homo sapiens |

BVYTAWGTAAACAAWV |
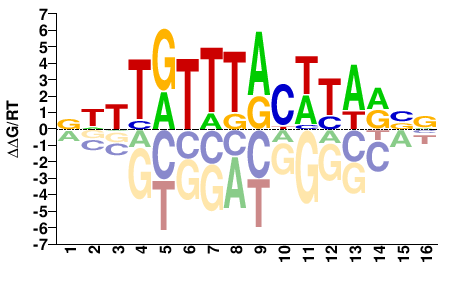
BWTTGTTTACWTARBV |
SELEX
Yin et al.(2017)
FOXA1_FL_Methyl-HT-SELEX_1
|
0.850
|
0.907
|
FOXA1
M04822_2.00 |
Homo sapiens |
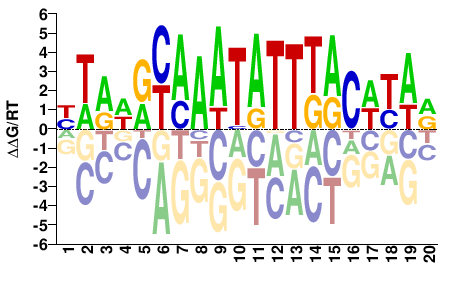
YTRWRYAAATATTTACWYAR |
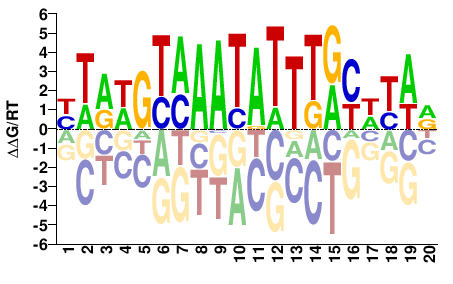
YTRWGTAAATATTTRYWYAR |
SELEX
Yin et al.(2017)
FOXA1_FL_Methyl-HT-SELEX_2
|
0.850
|
0.907
|
Foxa2
M00162_2.00 |
Mus musculus |

GTAAAYAW |

WTRTTTAC |
PBM
Badis et al.(2009)
Foxa2_2830
|
0.849
|
0.930
|
FOXA2
M04815_2.00 |
Homo sapiens |
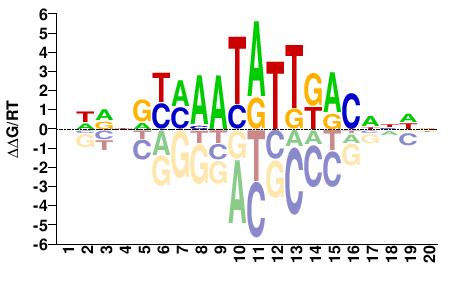
NHVNRYMAATATTKACNNDN |
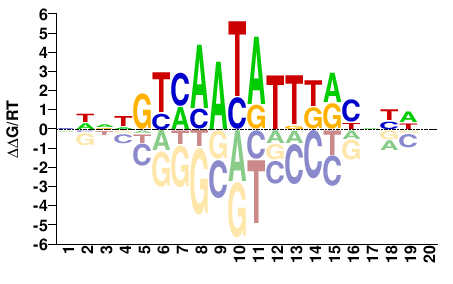
NHNNGTMAATATTKRYNBDN |
SELEX
Yin et al.(2017)
FOXA2_eDBD_HT-SELEX
|
0.849
|
0.930
|
Foxa2
M08121_2.00 |
Mus musculus |

TRTTTACWYWDN |

NHWRWGTAAAYA |
ChIP-seq
Mathelier et al.(2014)
MA0047.2
|
0.849
|
0.930
|
FOXA2
M07959_2.00 |
Homo sapiens |

NNWRWGYAAAYANNN |
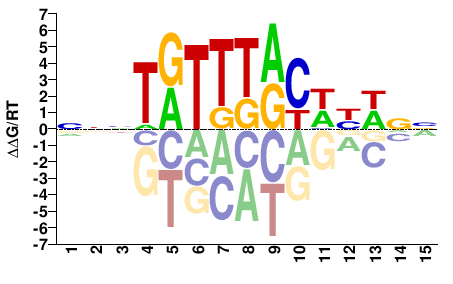
NNNTRTTTRCWYWNN |
ChIP-seq
Gerstein et al.(2012)
HepG2_FOXA2_HudsonAlpha
|
0.849
|
0.930
|
FOXA2
M09088_2.00 |
Homo sapiens |

NHWRWGTAAACA |

TGTTTACWYWDN |
Misc
Kulakovskiy et al.(2013)
FOXA2_HUMAN.H11MO.0.A
|
0.849
|
0.930
|
Foxa2
M09104_2.00 |
Mus musculus |
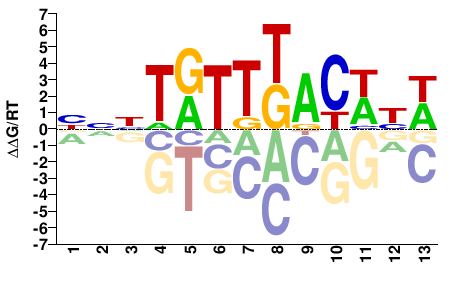
BNHTRTTKACWYW |
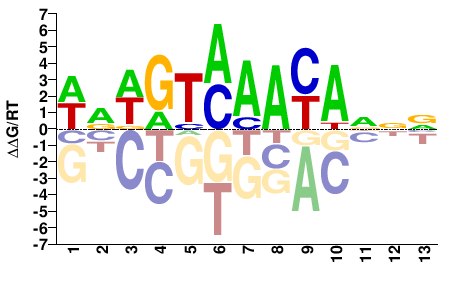
WRWGTMAAYADNV |
Misc
Kulakovskiy et al.(2013)
FOXA2_MOUSE.H11MO.0.A
|
0.849
|
0.930
|
FOXA2
M09542_2.00 |
Homo sapiens |

WVWGTMAACANV |

BNTGTTKACWBW |
Misc
Heinz et al.(2010)
Liver-Foxa2_GSE25694
|
0.849
|
0.930
|
FOXA2
M10540_2.00 |
Homo sapiens |
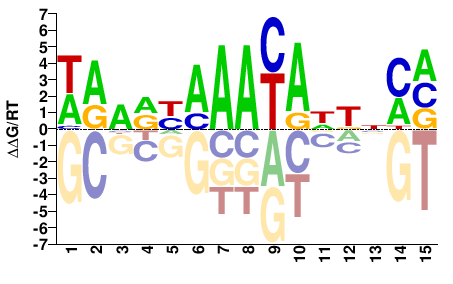
WAARYAAAYADKNMV |
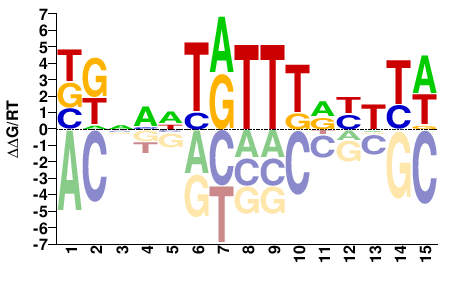
BKNMHTRTTTRYTTW |
Transfac
Matys et al.(2006)
V$HNF3B_01
|
0.849
|
0.930
|
FOXA2
M10541_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$HNF3B_Q6
|
0.849
|
0.930
|
AAA20679
M10575_2.00 |
Xenopus laevis |

WNWGTMAACAWWMW |

WKWWTGTTKACWNW |
Transfac
Matys et al.(2006)
V$XFD3_01
|
0.849
|
0.930
|
FOXA2
M04816_2.00 |
Homo sapiens |

NWRNRYMAATATTKACWNWN |

NWNWGTMAATATTKRYNYWN |
SELEX
Yin et al.(2017)
FOXA2_eDBD_Methyl-HT-SELEX
|
0.849
|
0.930
|
Q2L6N7_PELSI
M01990_2.00 |
Trionyx sinensis |

KTRTTTACA |

TGTAAAYAM |
PBM
Weirauch et al.(2014)
pTH5334
|
0.837
|
0.849
|
FOXA3
M04831_2.00 |
Homo sapiens |

BVYTAWGTAAACAAAN |

NTTTGTTTACWTARBV |
SELEX
Yin et al.(2017)
FOXA3_FL_HT-SELEX_1
|
0.817
|
0.872
|
FOXA3
M04832_2.00 |
Homo sapiens |

HWRWGYAAATATTKACWYWD |

HWRWGTMAATATTTRCWYWD |
SELEX
Yin et al.(2017)
FOXA3_FL_HT-SELEX_2
|
0.817
|
0.872
|
FOXA3
M09096_2.00 |
Homo sapiens |

HWRWGTMAAYADN |

NHTRTTKACWYWD |
Misc
Kulakovskiy et al.(2013)
FOXA3_HUMAN.H11MO.0.B
|
0.817
|
0.872
|
Foxa3
M09106_2.00 |
Mus musculus |

HWRWGTMAAYADN |

NHTRTTKACWYWD |
Misc
Kulakovskiy et al.(2013)
FOXA3_MOUSE.H11MO.0.A
|
0.817
|
0.872
|
FOXA3
M10557_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$HNF3G_Q4
|
0.817
|
0.872
|
FOXA3
M04833_2.00 |
Homo sapiens |

CVYTAWGTAAACAAWB |
VWTTGTTTACWTARBG |
SELEX
Yin et al.(2017)
FOXA3_FL_Methyl-HT-SELEX_1
|
0.817
|
0.872
|
FOXA3
M04834_2.00 |
Homo sapiens |
HWRWGYAAATATTKACWYWD |

HWRWGTMAATATTTRCWYWD |
SELEX
Yin et al.(2017)
FOXA3_FL_Methyl-HT-SELEX_2
|
0.817
|
0.872
|
pha-4
M08123_2.00 |
Caenorhabditis elegans |

WRWGYAAAYA |

TRTTTRCWYW |
ChIP-seq
Mathelier et al.(2014)
MA0546.1
|
0.780
|
0.837
|
pha-4
M08201_2.00 |
Caenorhabditis elegans |

GTAAACAR |
YTGTTTAC |
ChIP-seq
Contrino et al.(2012)
Mw164
|
0.780
|
0.837
|
pha-4
M09548_2.00 |
Caenorhabditis elegans |

RYAMAYAN |

NTRTKTRY |
Misc
Heinz et al.(2010)
cElegans-Embryos-PHA4_modEncode
|
0.780
|
0.837
|
Foxb1
M01983_2.00 |
Mus musculus |

TRTTKAYWYW |

WRWRTMAAYA |
PBM
Weirauch et al.(2014)
pTH2808
|
0.723
|
0.767
|
FOXB1
M03028_2.00 |
Homo sapiens |
GAATGACACRGCKV |
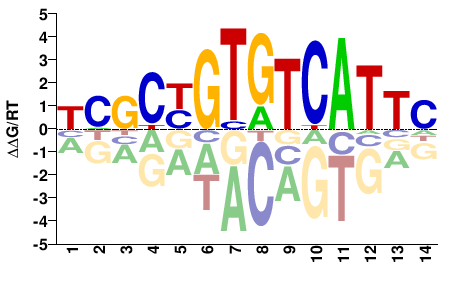
BMGCYGTGTCATTC |
SELEX
Jolma et al.(2013)
FOXB1_1
|
0.723
|
0.767
|
FOXB1
M03029_2.00 |
Homo sapiens |
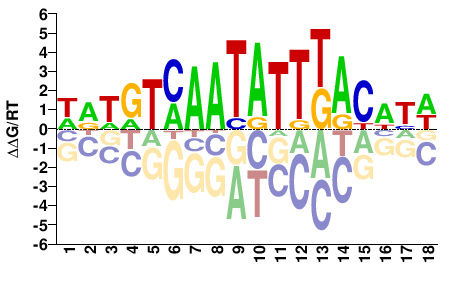
WRWGTMAATATTKACWYW |
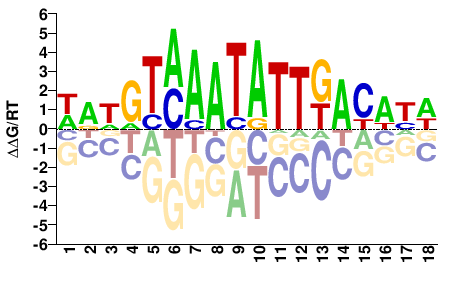
WRWGTMAATATTKACWYW |
SELEX
Jolma et al.(2013)
FOXB1_2
|
0.723
|
0.767
|
FOXB1
M03030_2.00 |
Homo sapiens |

WTRTTKACWTW |

WAWGTMAAYAW |
SELEX
Jolma et al.(2013)
FOXB1_3
|
0.723
|
0.767
|
FOXB1
M03031_2.00 |
Homo sapiens |
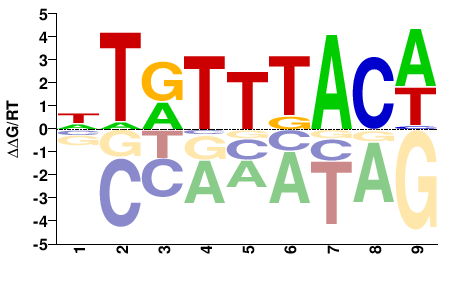

NTRTTTACW |
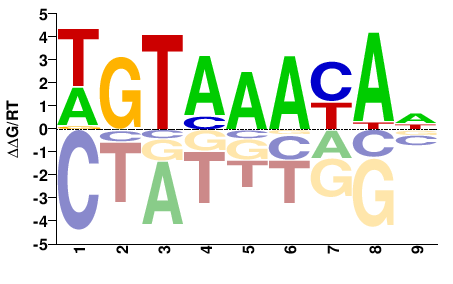

WGTAAAYAN |
SELEX
Jolma et al.(2013)
FOXB1_4
|
0.723
|
0.767
|
FOXB1
M04837_2.00 |
Homo sapiens |

NWRDGYMAATATTKRCWYWD |
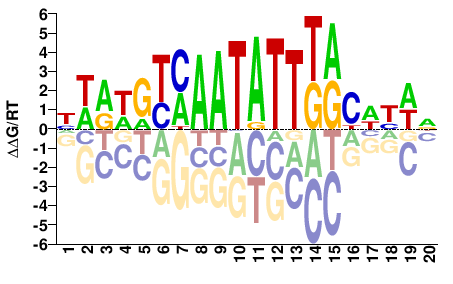
HWRWGYMAATATTKRCHYWN |
SELEX
Yin et al.(2017)
FOXB1_eDBD_HT-SELEX
|
0.723
|
0.767
|
FOXB1
M04838_2.00 |
Homo sapiens |

NWRNGYMAATATTKRCDYWD |
HWRHGYMAATATTKRCNYWN |
SELEX
Yin et al.(2017)
FOXB1_eDBD_Methyl-HT-SELEX
|
0.723
|
0.767
|
fd96Ca
M03743_2.00 |
Drosophila melanogaster |

WRWGYMAATATTKRCWYW |
WRWGYMAATATTKRCWYW |
SELEX
Nitta et al.(2015)
fd96Ca_1
|
0.715
|
0.709
|
fd64A
M06197_2.00 |
Drosophila melanogaster |

KVWAAACA |

TGTTTWBM |
B1H
Zhu et al.(2011)
fd64A_SANGER_5_FBgn0004895
|
0.674
|
0.628
|
FOXC1
M00257_2.00 |
Homo sapiens |
NNNVHHAAYA |
TRTTDDBNNN |
PBM
Barrera et al.(2016)
FOXC1_REF
|
0.665
|
0.756
|
Foxc2
M00796_2.00 |
Mus musculus |
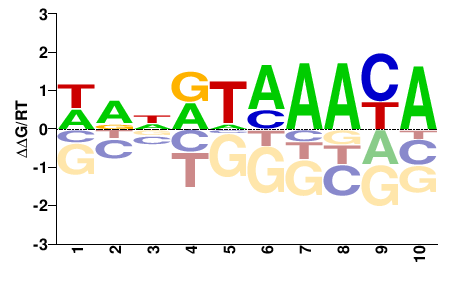
HNNRHMAAYA |
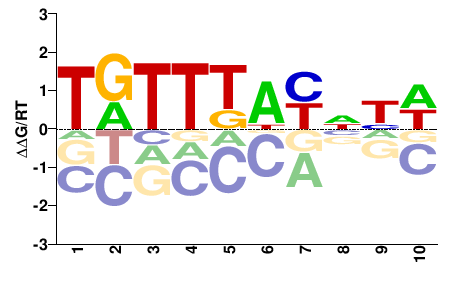
TRTTKDYNND |
PBM
Weirauch et al.(2013)
pTH3796
|
0.665
|
0.756
|
Foxc1
M01982_2.00 |
Mus musculus |

HNNRHMAAYA |

TRTTKDYNND |
PBM
Weirauch et al.(2014)
pTH2673
|
0.665
|
0.756
|
FOXC1
M03013_2.00 |
Homo sapiens |

AWRTAAAYAAAYA |

TRTTTRTTTAYWT |
SELEX
Jolma et al.(2013)
FOXC1_1
|
0.665
|
0.756
|
FOXC1
M03014_2.00 |
Homo sapiens |

NRWMAATATTKWYN |

NRWMAATATTKWYN |
SELEX
Jolma et al.(2013)
FOXC1_2
|
0.665
|
0.756
|
FOXC1
M03015_2.00 |
Homo sapiens |

WRWRTMAAYAW |

WTRTTKAYWYW |
SELEX
Jolma et al.(2013)
FOXC1_3
|
0.665
|
0.756
|
FOXC2
M03036_2.00 |
Homo sapiens |

NRTMAATATTDWYN |

NRWHAATATTKAYN |
SELEX
Jolma et al.(2013)
FOXC2_1
|
0.665
|
0.756
|
FOXC2
M03037_2.00 |
Homo sapiens |

RTAAAYAAACA |

TGTTTRTTTAY |
SELEX
Jolma et al.(2013)
FOXC2_2
|
0.665
|
0.756
|
FOXC2
M03038_2.00 |
Homo sapiens |
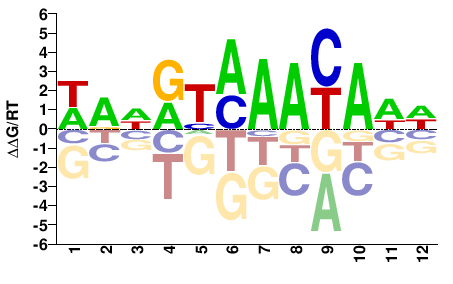
WAHRTMAAYAWW |

WWTRTTKAYDTW |
SELEX
Jolma et al.(2013)
FOXC2_3
|
0.665
|
0.756
|
Foxc1
M03059_2.00 |
Mus musculus |

RTAAAYAAACA |

TGTTTRTTTAY |
SELEX
Jolma et al.(2013)
Foxc1_1
|
0.665
|
0.756
|
Foxc1
M03060_2.00 |
Mus musculus |

RTAAACA |

TGTTTAY |
SELEX
Jolma et al.(2013)
Foxc1_2
|
0.665
|
0.756
|
FOXC2
M04841_2.00 |
Homo sapiens |
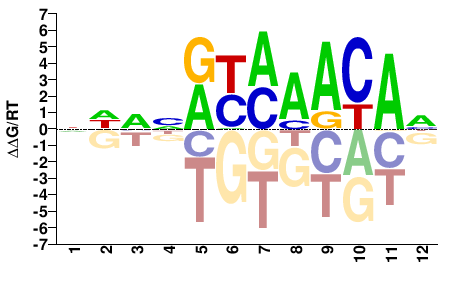
NHVNRYMAACAH |

DTGTTKRYNBDN |
SELEX
Yin et al.(2017)
FOXC2_eDBD_HT-SELEX_1
|
0.665
|
0.756
|
FOXC2
M04842_2.00 |
Homo sapiens |

RHAMAYAAACAH |
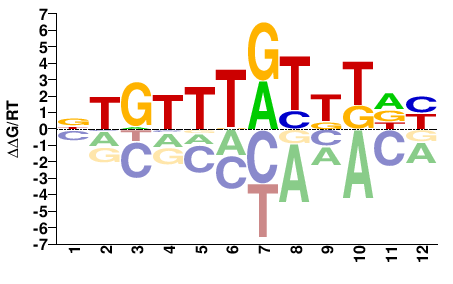
DTGTTTRTKTDY |
SELEX
Yin et al.(2017)
FOXC2_eDBD_HT-SELEX_2
|
0.665
|
0.756
|
FOXC1
M09083_2.00 |
Homo sapiens |
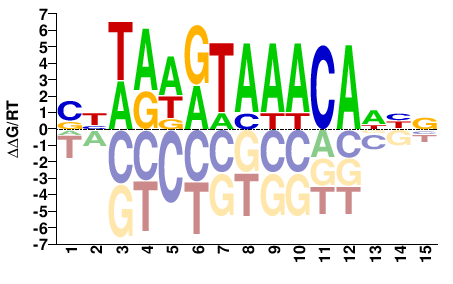
SBWAWRTAAACADHN |

NDHTGTTTAYWTWVS |
Misc
Kulakovskiy et al.(2013)
FOXC1_HUMAN.H11MO.0.C
|
0.665
|
0.756
|
Foxc1
M09111_2.00 |
Mus musculus |
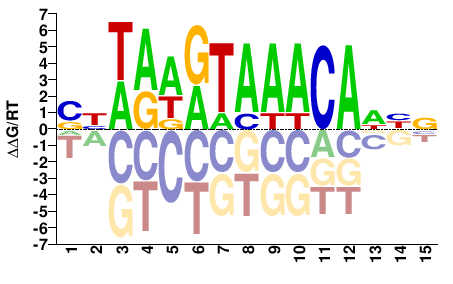
SBWAWRTAAACADHN |

NDHTGTTTAYWTWVS |
Misc
Kulakovskiy et al.(2013)
FOXC1_MOUSE.H11MO.0.C
|
0.665
|
0.756
|
FOXC2
M10559_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$FKHL14_Q2
|
0.665
|
0.756
|
FOXC1
M10525_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$FOXC1_Q4
|
0.665
|
0.756
|
FOXC1
M10526_2.00 |
Homo sapiens |
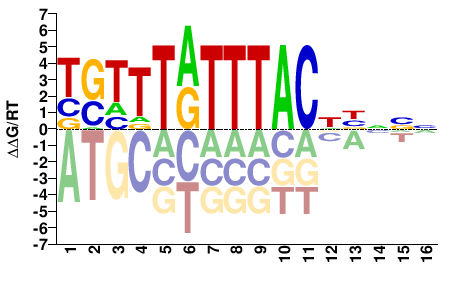
TSTTTATTTACDBNVN |

NBNVHGTAAATAAASA |
Transfac
Matys et al.(2006)
V$FREAC3_01
|
0.665
|
0.756
|
FOXC2
M04843_2.00 |
Homo sapiens |
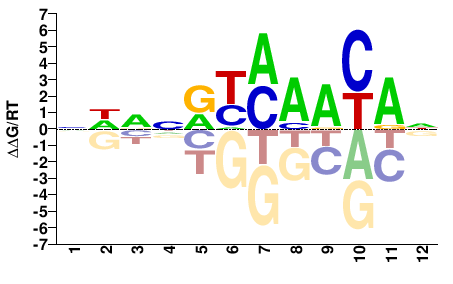
NHNNRYMAACAN |
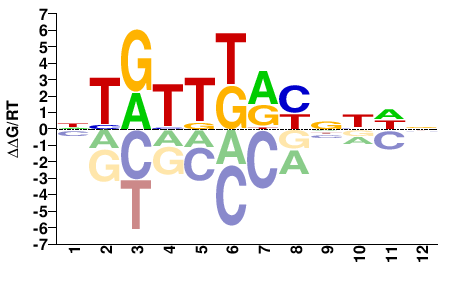
NTGTTKRYNNDN |
SELEX
Yin et al.(2017)
FOXC2_eDBD_Methyl-HT-SELEX_1
|
0.665
|
0.756
|
FOXC2
M04844_2.00 |
Homo sapiens |

VHMMAYAAACAN |

NTGTTTRTKKDB |
SELEX
Yin et al.(2017)
FOXC2_eDBD_Methyl-HT-SELEX_2
|
0.665
|
0.756
|
FOXC1
M00253_2.00 |
Homo sapiens |
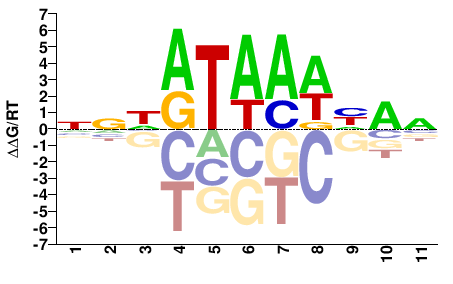
NNHATAAWHAN |
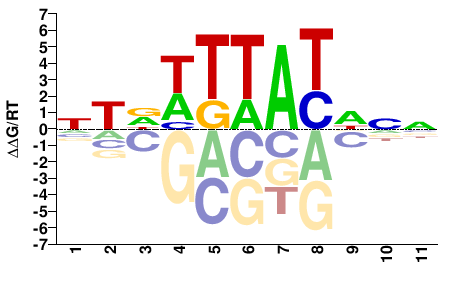
NTDWTTATDNN |
PBM
Barrera et al.(2016)
FOXC1_F112S
|
0.665
|
0.744
|
FOXC1
M00255_2.00 |
Homo sapiens |
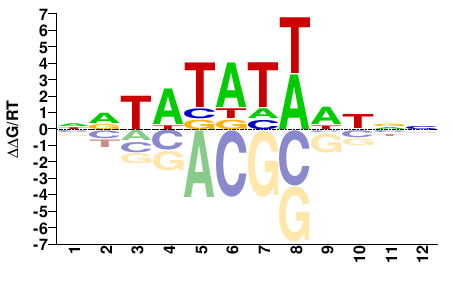

NRTATATWHNNN |
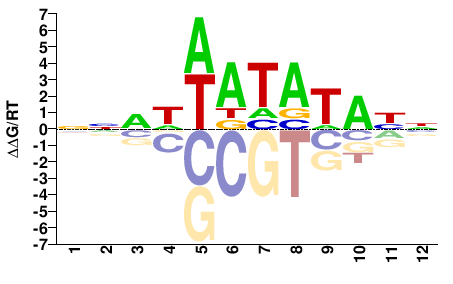
NNNDWATATAYN |
PBM
Barrera et al.(2016)
FOXC1_L130F
|
0.665
|
0.744
|
FOXC1
M00256_2.00 |
Homo sapiens |
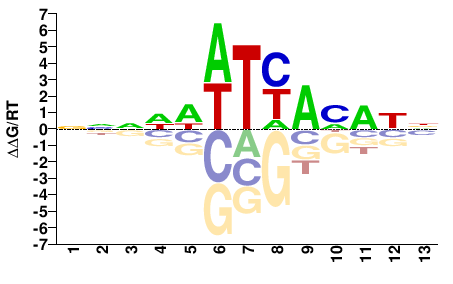
NNNNWWTYAMAWN |
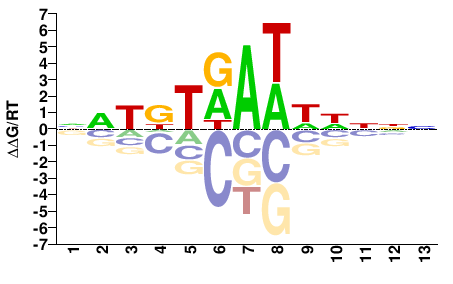
NWTKTRAWWNNNN |
PBM
Barrera et al.(2016)
FOXC1_P79L
|
0.665
|
0.744
|
FOXC1
M00258_2.00 |
Homo sapiens |
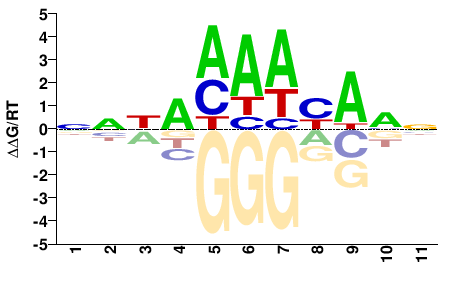
NNNAMAAYANN |
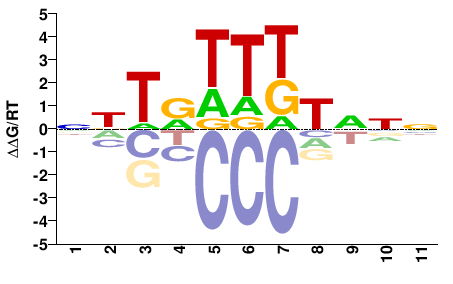

NNTRTTKTNNN |
PBM
Barrera et al.(2016)
FOXC1_S131L
|
0.665
|
0.744
|
FOXC1
M00259_2.00 |
Homo sapiens |

RTAAACAW |

WTGTTTAY |
PBM
Barrera et al.(2016)
FOXC1_S82T
|
0.665
|
0.744
|
| For this family, TFs with SR scores > 0.664 will likely have a similar motif |