|
|
ESRRG
(Homo sapiens)
Nuclear receptor
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
Animal TF db |
| PF00105 (zf-C4) |
IPR001628 |
ENSG00000196482 |
T303248_2.00 |
Ensembl (2018-Dec-8) |
Link out |
| NCBI Gene Info:This gene encodes a member of the estrogen receptor-related receptor (ESRR) family, which belongs to the nuclear hormone receptor superfamily. All members of the ESRR family share an almost identical DNA binding domain, which is composed of two C4-type zinc finger motifs. The ESRR members are orphan nuclear receptors; they bind to the estrogen response element and steroidogenic factor 1 response element, and activate genes controlled by both response elements in the absence of any ligands. The ESRR family is closely related to the estrogen receptor (ER) family. They share target genes, co-regulators and promoters, and by targeting the same set of genes, the ESRRs seem to interfere with the ER-mediated estrogen response in various ways. It has been reported that the family member encoded by this gene functions as a transcriptional activator of DNA cytosine-5-methyltransferases 1 (Dnmt1) expression by direct binding to its response elements in the DNMT1 promoters, modulates cell proliferation and estrogen signaling in breast cancer, and negatively regulates bone morphogenetic protein 2-induced osteoblast differentiation and bone formation. Multiple alternatively spliced transcript variants have been identified, which mainly differ at the 5' end and some of which encode protein isoforms differing in the N-terminal region. [provided by RefSeq, Aug 2011] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
ESRRG
M03412_2.00 |
Homo sapiens |
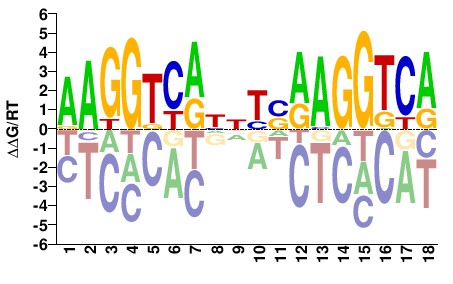
AAGGTCAHNYSAAGGTCA |
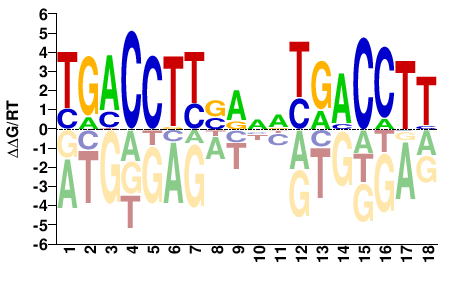
TGACCTTSRNDTGACCTT |
SELEX
Jolma et al.(2013)
ESRRG_1 |
(Direct) |
(Direct) |
ESRRG
M03413_2.00 |
Homo sapiens |
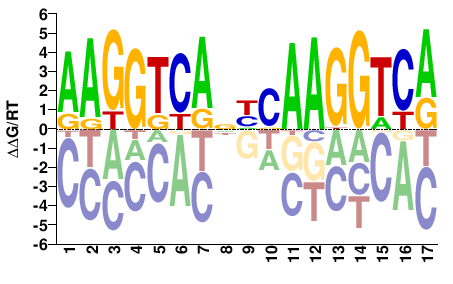
AAGGTCANHCAAGGTCA |
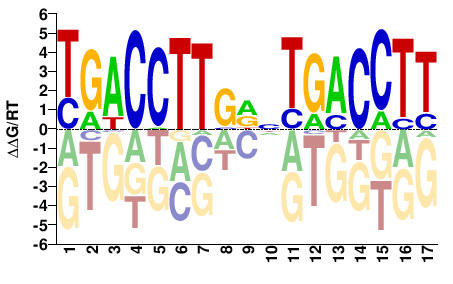
TGACCTTGDNTGACCTT |
SELEX
Jolma et al.(2013)
ESRRG_2 |
(Direct) |
(Direct) |
ESRRG
M03414_2.00 |
Homo sapiens |

YSAAGGTCAH |

DTGACCTTSR |
SELEX
Jolma et al.(2013)
ESRRG_3 |
(Direct) |
(Direct) |
ESRRG
M05673_2.00 |
Homo sapiens |
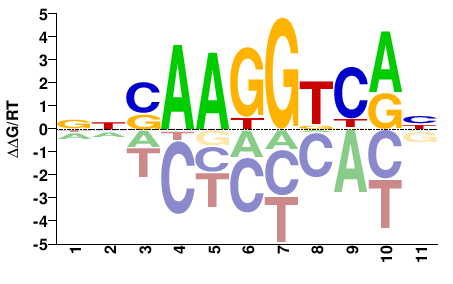
NNSAAGGTCRN |

NYGACCTTSNN |
SELEX
Yin et al.(2017)
ESRRG_eDBD_HT-SELEX |
(Direct) |
(Direct) |
ESRRG
M05674_2.00 |
Homo sapiens |
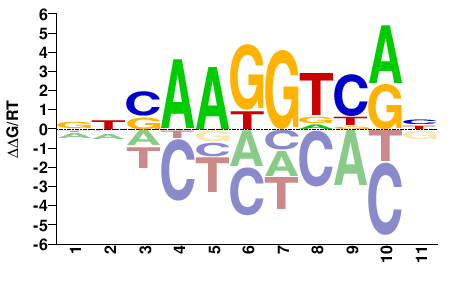
NNSAAGGTCRN |

NYGACCTTSNN |
SELEX
Yin et al.(2017)
ESRRG_eDBD_Methyl-HT-SELEX |
(Direct) |
(Direct) |
ESRRG
M05675_2.00 |
Homo sapiens |
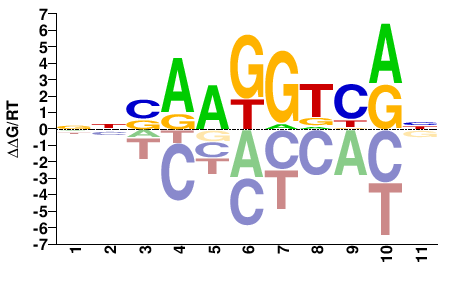
NNSAAGGTCRN |

NYGACCTTSNN |
SELEX
Yin et al.(2017)
ESRRG_FL_HT-SELEX |
(Direct) |
(Direct) |
ESRRG
M05676_2.00 |
Homo sapiens |

NNSAAGGTCRN |

NYGACCTTSNN |
SELEX
Yin et al.(2017)
ESRRG_FL_Methyl-HT-SELEX |
(Direct) |
(Direct) |
ESRRG
M09296_2.00 |
Homo sapiens |

TCAAGGTCA |

TGACCTTGA |
Misc
Kulakovskiy et al.(2013)
ERR3_HUMAN.H11MO.0.B |
(Direct) |
(Direct) |
ESRRG
M11185_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERR3_Q2_01 |
(Direct) |
(Direct) |
ESRRG
M11186_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERR3_Q2 |
(Direct) |
(Direct) |
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
Esrrg
M00817_2.00 |
Mus musculus |

VAAGGTCA |

TGACCTTB |
PBM
Weirauch et al.(2013)
pTH3841
|
0.985
|
1.000
|
ESRRB
M03367_2.00 |
Homo sapiens |

TCAAGGTCAWH |

DWTGACCTTGA |
SELEX
Jolma et al.(2013)
ESRRB_1
|
0.985
|
1.000
|
ESRRB
M05595_2.00 |
Homo sapiens |

BBSAAGGTCRH |

DYGACCTTSVV |
SELEX
Yin et al.(2017)
ESRRB_eDBD_HT-SELEX
|
0.985
|
1.000
|
ESRRB
M09272_2.00 |
Homo sapiens |

BCAAGGTCA |

TGACCTTGV |
Misc
Kulakovskiy et al.(2013)
ERR2_HUMAN.H11MO.0.A
|
0.985
|
1.000
|
Esrrg
M09321_2.00 |
Mus musculus |

BSAAGGWCAB |

VTGWCCTTSV |
Misc
Kulakovskiy et al.(2013)
ERR3_MOUSE.H11MO.0.C
|
0.985
|
1.000
|
ESRRB
M11129_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERR2_01
|
0.985
|
1.000
|
ESRRB
M11130_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERR2_Q2
|
0.985
|
1.000
|
ESRRB
M11131_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERR2_Q3
|
0.985
|
1.000
|
ESRRB
M05596_2.00 |
Homo sapiens |
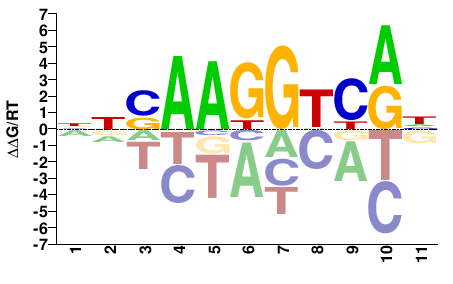
NNSAAGGTCRH |

DYGACCTTSNN |
SELEX
Yin et al.(2017)
ESRRB_eDBD_Methyl-HT-SELEX
|
0.985
|
1.000
|
Esrrb
M08034_2.00 |
Mus musculus |

NNBYCAAGGTCANNB |
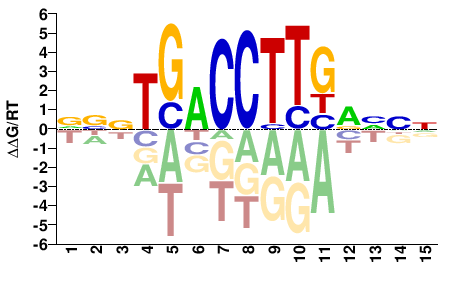
VNNTGACCTTGRVNN |
ChIP-seq
Chen et al.(2011)
SRP000217_Esrrb
|
0.985
|
0.986
|
Esrrb
M09312_2.00 |
Mus musculus |

NHDBTCAAGGTCANNBN |
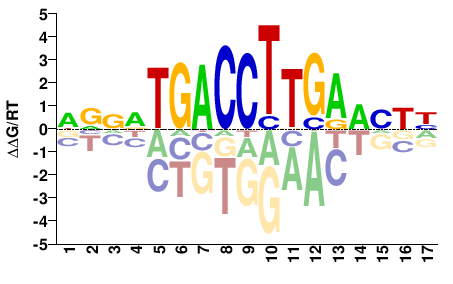
NVNNTGACCTTGAVHDN |
Misc
Kulakovskiy et al.(2013)
ERR2_MOUSE.H11MO.0.A
|
0.985
|
0.986
|
Esrra
M00183_2.00 |
Mus musculus |

VAAGGTCAH |
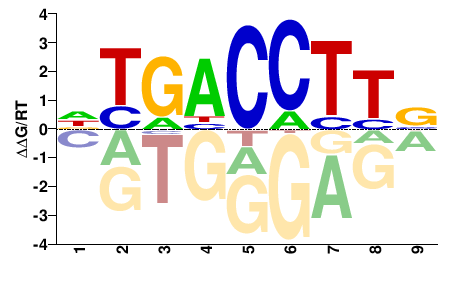
DTGACCTTB |
PBM
Badis et al.(2009)
Esrra_2190
|
0.976
|
0.914
|
ESRRA
M03397_2.00 |
Homo sapiens |

NYSAAGGTCAH |

DTGACCTTSRN |
SELEX
Jolma et al.(2013)
ESRRA_1
|
0.976
|
0.914
|
ESRRA
M03398_2.00 |
Homo sapiens |

AAGGTCAWHCAAGGTCA |
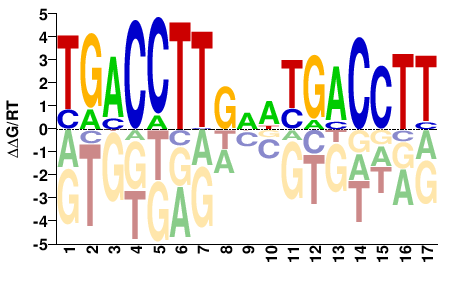
TGACCTTGDWTGACCTT |
SELEX
Jolma et al.(2013)
ESRRA_2
|
0.976
|
0.914
|
ESRRA
M03399_2.00 |
Homo sapiens |
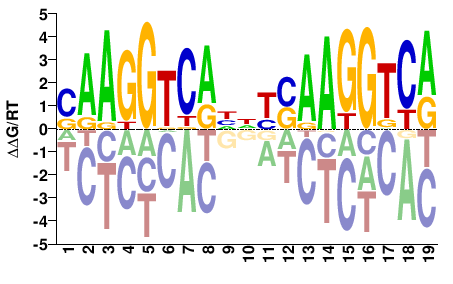
SAAGGTCAHNYSAAGGTCA |
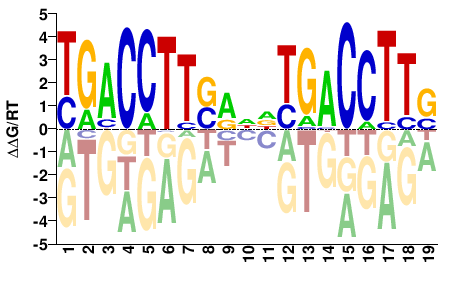
TGACCTTSRNDTGACCTTS |
SELEX
Jolma et al.(2013)
ESRRA_3
|
0.976
|
0.914
|
ESRRA
M03400_2.00 |
Homo sapiens |

NYSAAGGTCAH |

DTGACCTTSRN |
SELEX
Jolma et al.(2013)
ESRRA_4
|
0.976
|
0.914
|
ESRRA
M03401_2.00 |
Homo sapiens |
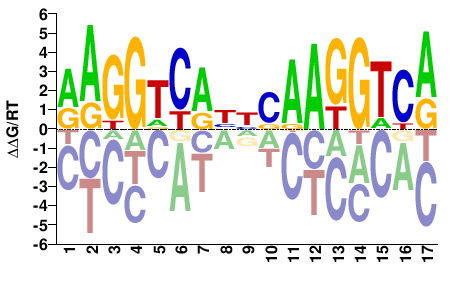
RAGGTCABNCAAGGTCA |
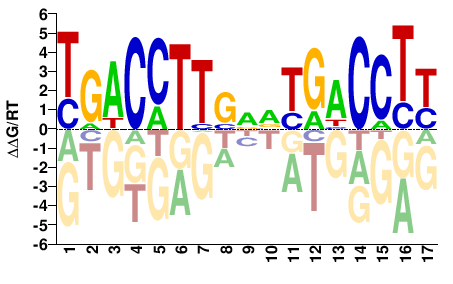
TGACCTTGNVTGACCTY |
SELEX
Jolma et al.(2013)
ESRRA_5
|
0.976
|
0.914
|
ESRRA
M03402_2.00 |
Homo sapiens |
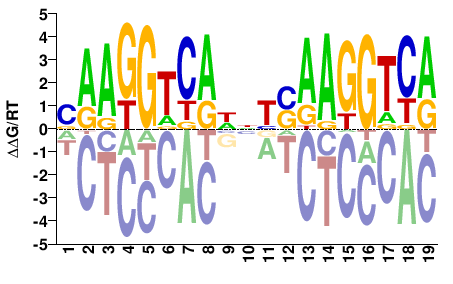
VAAGGTCAHNYSAAGGTCA |
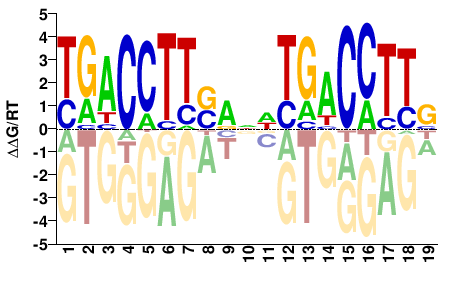
TGACCTTSRNDTGACCTTB |
SELEX
Jolma et al.(2013)
ESRRA_6
|
0.976
|
0.914
|
Esrra
M03430_2.00 |
Mus musculus |

DAGGTYAKTMAAGGTCA |

TGACCTTKAMTRACCTH |
SELEX
Jolma et al.(2013)
Esrra_1
|
0.976
|
0.914
|
Esrra
M03431_2.00 |
Mus musculus |

KTCAAGGTCAT |

ATGACCTTGAM |
SELEX
Jolma et al.(2013)
Esrra_2
|
0.976
|
0.914
|
ESRRA
M05655_2.00 |
Homo sapiens |
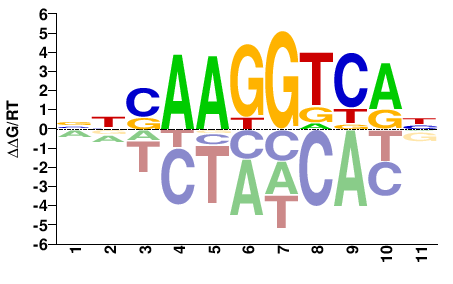
NNSAAGGTCAN |

NTGACCTTSNN |
SELEX
Yin et al.(2017)
ESRRA_eDBD_HT-SELEX
|
0.976
|
0.914
|
ESRRA
M05657_2.00 |
Homo sapiens |
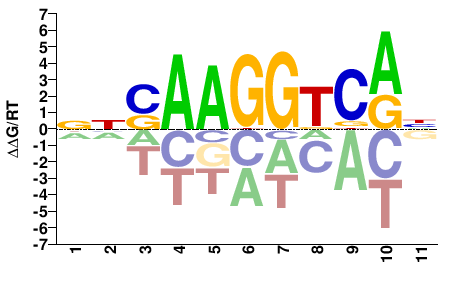
NNSAAGGTCAH |

DTGACCTTSNN |
SELEX
Yin et al.(2017)
ESRRA_FL_HT-SELEX
|
0.976
|
0.914
|
ESRRA
M07988_2.00 |
Homo sapiens |

RGGKCASMNTGMCCY |

RGGKCANKSTGMCCY |
ChIP-seq
Gerstein et al.(2012)
ECC-1_ERALPHA_HudsonAlpha
|
0.976
|
0.914
|
ESRRA
M07989_2.00 |
Homo sapiens |
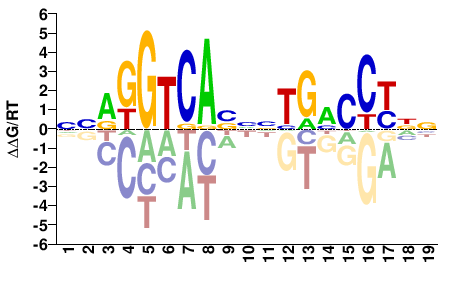

NNRGGTCABNNTGHCCYNN |
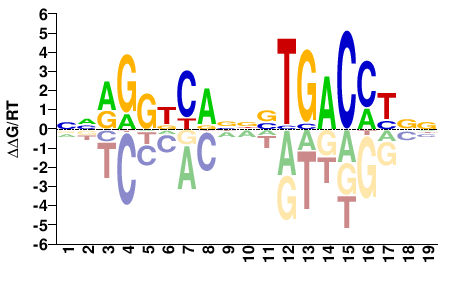

NNRGGDCANNVTGACCYNN |
ChIP-seq
Gerstein et al.(2012)
T-47D_ERALPHA_HudsonAlpha
|
0.976
|
0.914
|
ESRRA
M09290_2.00 |
Homo sapiens |
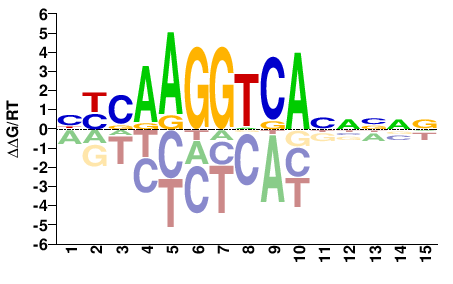
BYSAAGGTCANNNNN |
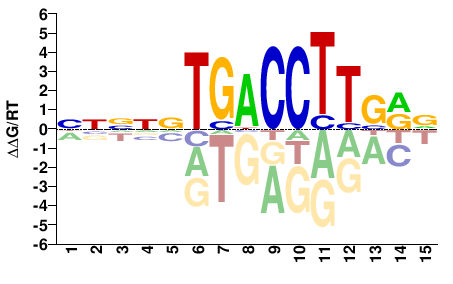
NNNNNTGACCTTSRV |
Misc
Kulakovskiy et al.(2013)
ERR1_HUMAN.H11MO.0.A
|
0.976
|
0.914
|
Esrra
M09319_2.00 |
Mus musculus |

NNBYCAAGGTCA |
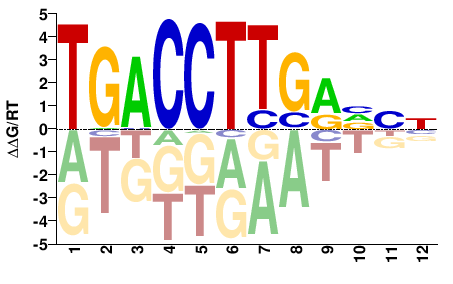

TGACCTTGRVNN |
Misc
Kulakovskiy et al.(2013)
ERR1_MOUSE.H11MO.0.A
|
0.976
|
0.914
|
ESRRA
M11168_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERR1_Q2_01
|
0.976
|
0.914
|
ESRRA
M11169_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERR1_Q2
|
0.976
|
0.914
|
ESRRA
M11170_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERR1_Q3
|
0.976
|
0.914
|
ESRRA
M11171_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ESRRA_10
|
0.976
|
0.914
|
ESRRA
M05656_2.00 |
Homo sapiens |
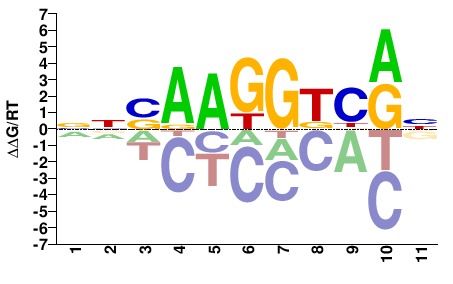
NNSAAGGTCRN |

NYGACCTTSNN |
SELEX
Yin et al.(2017)
ESRRA_eDBD_Methyl-HT-SELEX
|
0.976
|
0.914
|
ESRRA
M05658_2.00 |
Homo sapiens |
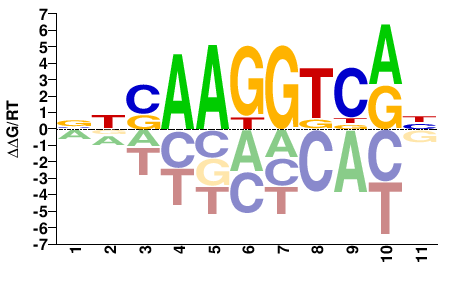
NBSAAGGTCRH |

DYGACCTTSVN |
SELEX
Yin et al.(2017)
ESRRA_FL_Methyl-HT-SELEX
|
0.976
|
0.914
|
ERR
M03941_2.00 |
Drosophila melanogaster |
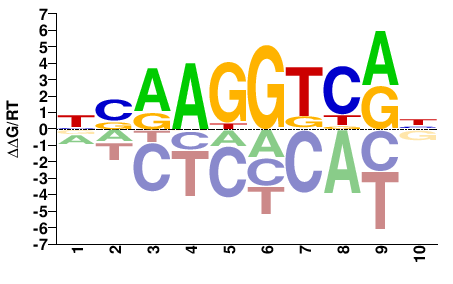
BSAAGGTCRH |

DYGACCTTSV |
SELEX
Nitta et al.(2015)
ERR_1
|
0.974
|
0.871
|
ERR
M06433_2.00 |
Drosophila melanogaster |

MAAGGTCA |

TGACCTTK |
B1H
Zhu et al.(2011)
ERR_SANGER_5_FBgn0035849
|
0.974
|
0.871
|
Esr1
M00813_2.00 |
Mus musculus |
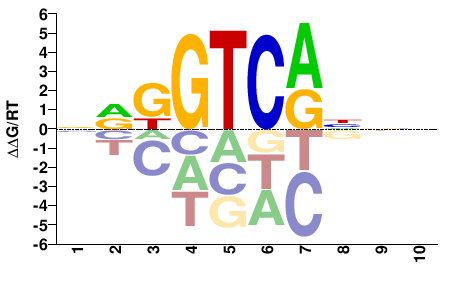
NRKGTCANNN |
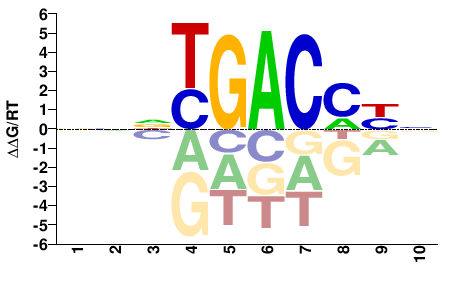
NNNTGACMYN |
PBM
Weirauch et al.(2013)
pTH3510
|
0.844
|
0.714
|
ESR1
M03356_2.00 |
Homo sapiens |
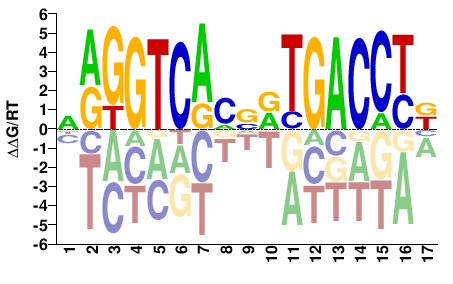
NRGGTCAMVRTGACCTK |
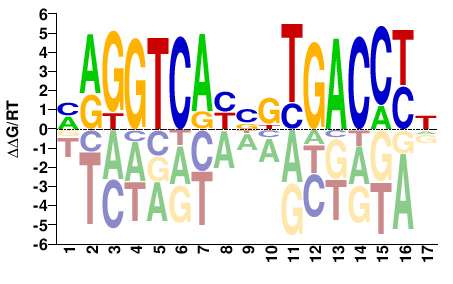

MAGGTCAYBKTGACCYN |
SELEX
Jolma et al.(2013)
ESR1_1
|
0.844
|
0.714
|
ESR1
M05569_2.00 |
Homo sapiens |
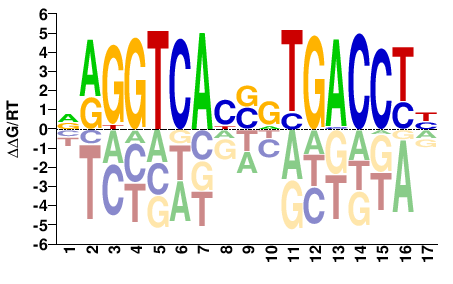

NAGGTCAYSRTGACCTN |
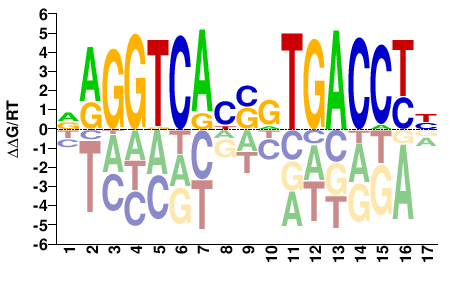
NAGGTCAYSRTGACCTN |
SELEX
Yin et al.(2017)
ESR1_eDBD_HT-SELEX
|
0.844
|
0.714
|
ESR1
M05571_2.00 |
Homo sapiens |
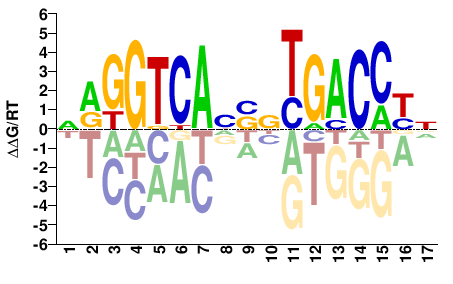
NRGGTCANSNTGACCYN |

NRGGTCANSNTGACCYN |
SELEX
Yin et al.(2017)
ESR1_FL_HT-SELEX
|
0.844
|
0.714
|
ESR1
M07985_2.00 |
Homo sapiens |

DNBYMAAGGTCANN |
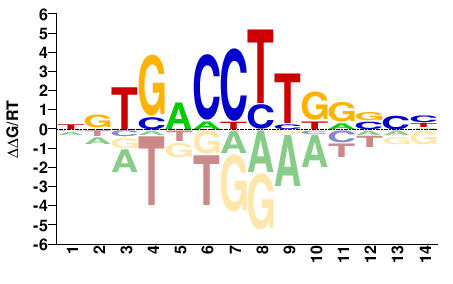
NNTGACCTTKRVNH |
ChIP-seq
Gerstein et al.(2012)
HepG2_ERRA_Stanford
|
0.844
|
0.714
|
ESR1
M08217_2.00 |
Homo sapiens |

NHNRGGTCANNNTGNCCY |

RGGNCANNNTGACCYNDN |
ChIP-seq
Contrino et al.(2012)
Mv66
|
0.844
|
0.714
|
ESR1
M08218_2.00 |
Homo sapiens |
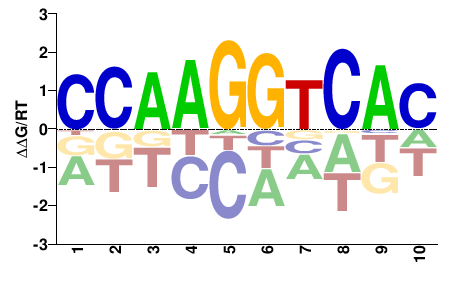

CCAAGGTCAS |

STGACCTTGG |
ChIP-seq
Contrino et al.(2012)
Mv67
|
0.844
|
0.714
|
ESR1
M09267_2.00 |
Homo sapiens |
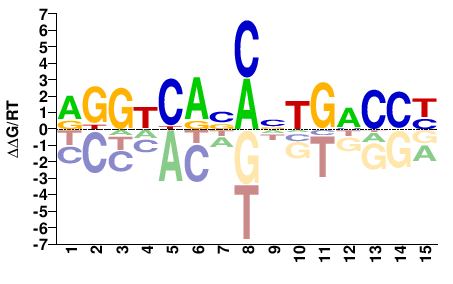
RGGKCABMNTGMCCY |

RGGKCANKVTGMCCY |
Misc
Kulakovskiy et al.(2013)
ESR1_HUMAN.H11MO.0.A
|
0.844
|
0.714
|
Esr1
M09309_2.00 |
Mus musculus |

RGGDCABKSTGWCCY |

RGGWCASMVTGHCCY |
Misc
Kulakovskiy et al.(2013)
ESR1_MOUSE.H11MO.0.A
|
0.844
|
0.714
|
ESR1
M09604_2.00 |
Homo sapiens |
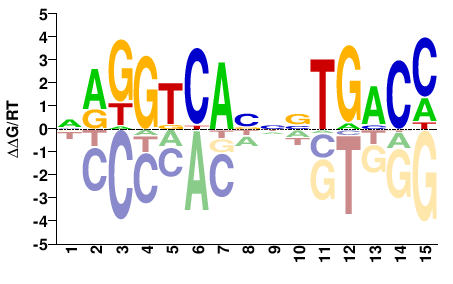
NRGGTCANNNTGACC |

GGTCANNNTGACCYN |
Misc
Heinz et al.(2010)
MCF7-ERa_Unpublished
|
0.844
|
0.714
|
ESR1
M11098_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERALPHA_01
|
0.844
|
0.714
|
ESR1
M11099_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERALPHA_Q4
|
0.844
|
0.714
|
ESR1
M11100_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERALPHA_Q6_01
|
0.844
|
0.714
|
ESR1
M11101_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERALPHA_Q6_02
|
0.844
|
0.714
|
ESR1
M11102_2.00 |
Homo sapiens |

NHRGGTCAVNNHNNHYBNN |
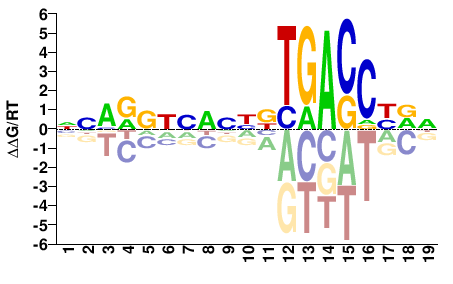
NNVRDNNDNNBTGACCYDN |
Transfac
Matys et al.(2006)
V$ER_Q6
|
0.844
|
0.714
|
ESR1
M11103_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ESR1_04
|
0.844
|
0.714
|
ESR1
M11104_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ESR1_05
|
0.844
|
0.714
|
ESR1
M05570_2.00 |
Homo sapiens |
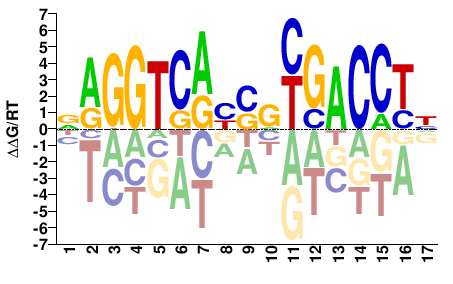

NAGGTCAYSRYGACCTH |
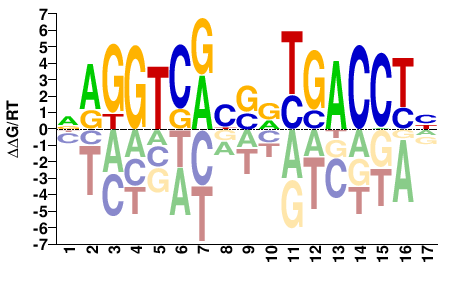
DAGGTCRYSRTGACCTN |
SELEX
Yin et al.(2017)
ESR1_eDBD_Methyl-HT-SELEX
|
0.844
|
0.714
|
ESR1
M05572_2.00 |
Homo sapiens |

NRGGTCANSBYGACCYN |
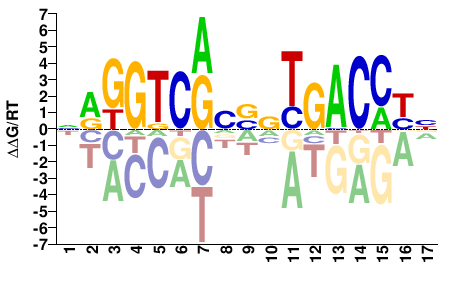
NRGGTCRVSNTGACCYN |
SELEX
Yin et al.(2017)
ESR1_FL_Methyl-HT-SELEX
|
0.844
|
0.714
|
Esr2
M01297_2.00 |
Mus musculus |
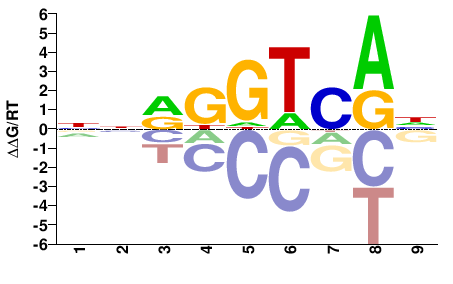
NNRGGTCAH |

DTGACCYNN |
PBM
Lambert et al.(2019)
pTH3511
|
0.839
|
0.700
|
ESR2
M08154_2.00 |
Homo sapiens |

RGGTCASNNWGNCCY |
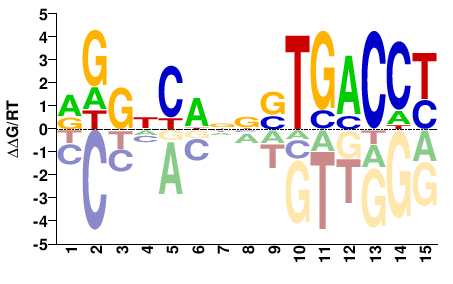

RGGNCWNNSTGACCY |
ChIP-seq
Mathelier et al.(2014)
MA0258.2
|
0.839
|
0.700
|
ESR2
M09279_2.00 |
Homo sapiens |

RGGKCRVRSYGMCCY |

RGGKCRSYBYGMCCY |
Misc
Kulakovskiy et al.(2013)
ESR2_HUMAN.H11MO.0.A
|
0.839
|
0.700
|
Esr2
M09311_2.00 |
Mus musculus |

RGGKCRVRSYGMCCY |

RGGKCRSYBYGMCCY |
Misc
Kulakovskiy et al.(2013)
ESR2_MOUSE.H11MO.0.A
|
0.839
|
0.700
|
ESR2
M11146_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERBETA_Q5_01
|
0.839
|
0.700
|
ESR2
M11147_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$ERBETA_Q5
|
0.839
|
0.700
|
Q6VU64_APLCA
M02423_2.00 |
Aplysia californica |

NRGGTCAN |
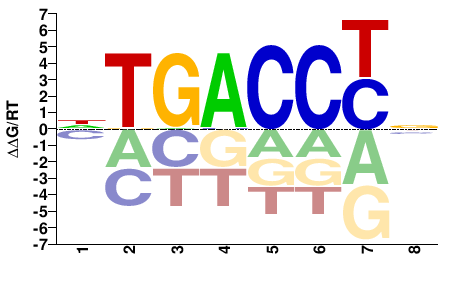
NTGACCYN |
PBM
Weirauch et al.(2014)
pTH6055
|
0.824
|
0.686
|
| For this family, TFs with SR scores > 0.745 will likely have a similar motif |