|
|
Tp53
(Rattus norvegicus)
p53
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
Animal TF db |
| PF00870 (P53) |
IPR011615 |
ENSRNOG00000010756 |
T311141_2.00 |
Ensembl (2018-Dec-8) |
Link out |
| NCBI Gene Info:This gene encodes tumor protein p53, which responds to diverse cellular stresses to regulate target genes that induce cell cycle arrest, apoptosis, senescence, DNA repair, or changes in metabolism. p53 protein is expressed at low level in normal cells and at a high level in a variety of transformed cell lines, where it is believed to contribute to transformation and malignancy. p53 is a DNA-binding protein containing transcription activation, DNA-binding, and oligomerization domains. It is postulated to bind to a p53-binding site and activate expression of downstream genes that inhibit growth and/or invasion, and thus function as a tumor suppressor. Alternatively spliced transcript variants have been found for this gene, but the biological validity of the variants has not been determined. p53 pseudogenes have been found on chromosomes 9 and 18. [provided by RefSeq, Jul 2008] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
| No direct experiments |
|
|
|
|
|
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
Trp53
M03439_2.00 |
Mus musculus |
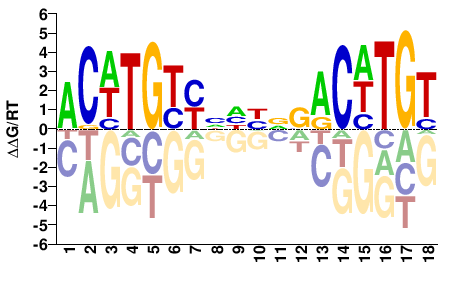
ACWTGYYNHHDSACWTGT |
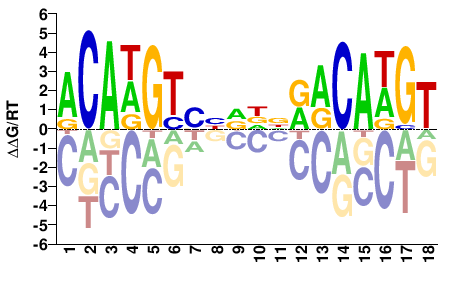
ACAWGTSHDDNRRCAWGT |
SELEX
Jolma et al.(2013)
Tp53_1
|
0.938
|
0.938
|
Trp53
M03440_2.00 |
Mus musculus |
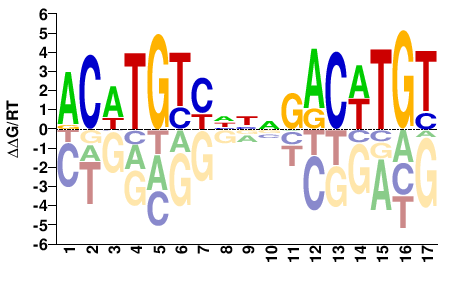
ACWTGTYHNNGACWTGT |
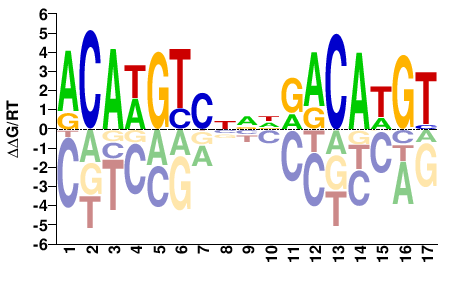
ACAWGTCNNDRACAWGT |
SELEX
Jolma et al.(2013)
Tp53_2
|
0.938
|
0.938
|
Trp53
M03441_2.00 |
Mus musculus |
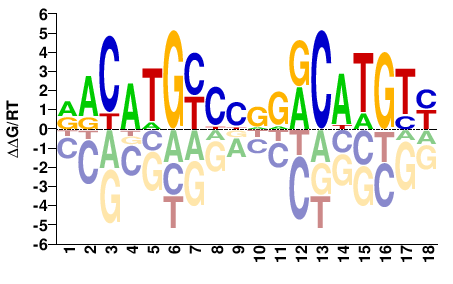
RACATGYCYRGRCATGTY |
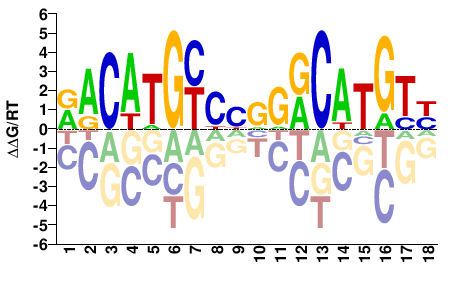
RACATGYCYRGRCATGTY |
SELEX
Jolma et al.(2013)
Tp53_3
|
0.871
|
0.871
|
Trp53
M09340_2.00 |
Mus musculus |
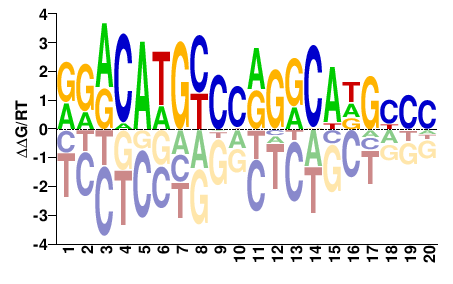
RRRCAWGYCCRGRCADGYMH |
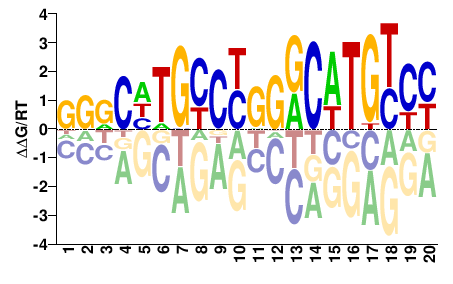

DKRCHTGYCYGGRCWTGYYY |
Misc
Kulakovskiy et al.(2013)
P53_MOUSE.H11MO.0.A
|
0.871
|
0.871
|
TP53
M09337_2.00 |
Homo sapiens |
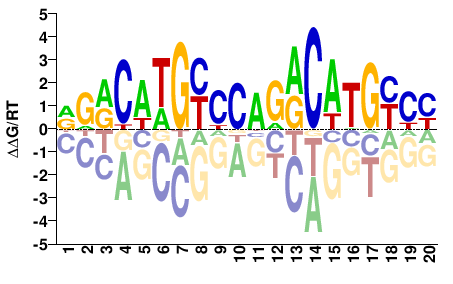
DRRCWTGYYCAGRCATGYYY |
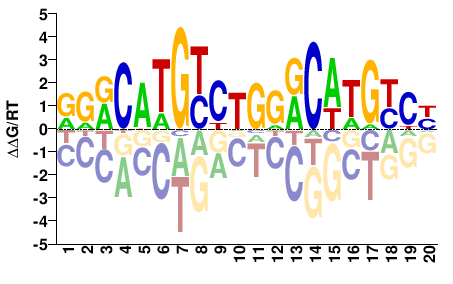
RRRCATGYCTGRRCAWGYYH |
Misc
Kulakovskiy et al.(2013)
P53_HUMAN.H11MO.0.A
|
0.848
|
0.848
|
TP53
M09621_2.00 |
Homo sapiens |
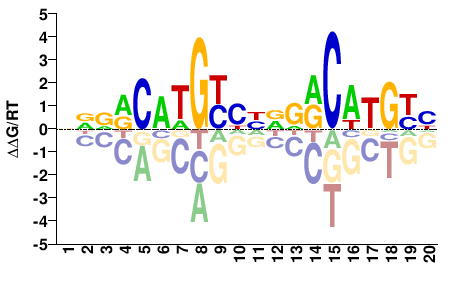
NNDRCAWGYHNNDRCWWGYH |
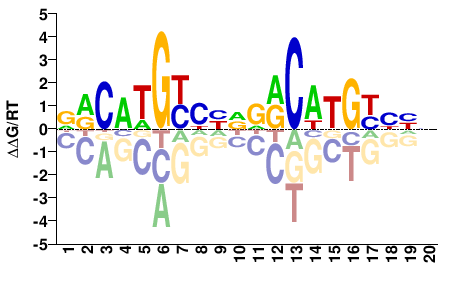

DRCWWGYHNNDRCWTGYHNN |
Misc
Heinz et al.(2010)
Saos-p53
|
0.848
|
0.848
|
TP53
M11197_2.00 |
Homo sapiens |

RGACATGCCCGGGCATGTYC |
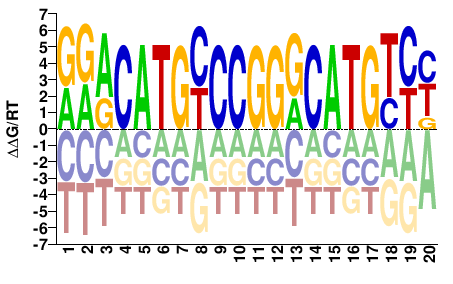
GRACATGCCCGGGCATGTCY |
Transfac
Matys et al.(2006)
V$P53_01
|
0.848
|
0.848
|
TP53
M11198_2.00 |
Homo sapiens |

NGRCWTGYYY |

RRRCAWGYCN |
Transfac
Matys et al.(2006)
V$P53_02
|
0.908
|
0.908
|
TP53
M11199_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$P53_03
|
0.908
|
0.908
|
TP53
M11200_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$P53_04
|
0.908
|
0.908
|
TP53
M11201_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$P53_05
|
0.848
|
0.848
|
TP53
M11202_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$P53_Q3_01
|
0.848
|
0.848
|
TP53
M11203_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$P53_Q3
|
0.848
|
0.848
|
| For this family, TFs with SR scores > 0.700 will likely have a similar motif |