|
|
SMAD1
(Homo sapiens)
SMAD
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
Animal TF db |
| PF03165 (MH1) |
IPR003619 |
ENSG00000170365 |
T324630_2.00 |
Ensembl (2018-Dec-8) |
Link out |
| NCBI Gene Info:The protein encoded by this gene belongs to the SMAD, a family of proteins similar to the gene products of the Drosophila gene 'mothers against decapentaplegic' (Mad) and the C. elegans gene Sma. SMAD proteins are signal transducers and transcriptional modulators that mediate multiple signaling pathways. This protein mediates the signals of the bone morphogenetic proteins (BMPs), which are involved in a range of biological activities including cell growth, apoptosis, morphogenesis, development and immune responses. In response to BMP ligands, this protein can be phosphorylated and activated by the BMP receptor kinase. The phosphorylated form of this protein forms a complex with SMAD4, which is important for its function in the transcription regulation. This protein is a target for SMAD-specific E3 ubiquitin ligases, such as SMURF1 and SMURF2, and undergoes ubiquitination and proteasome-mediated degradation. Alternatively spliced transcript variants encoding the same protein have been observed. [provided by RefSeq, Jul 2008] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
SMAD1
M11286_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SMAD1_01 |
(Direct) |
(Direct) |
SMAD1
M11287_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SMAD1_Q6 |
(Direct) |
(Direct) |
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
Smad5
M11288_2.00 |
Mus musculus |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SMAD5_01
|
0.952
|
0.952
|
SMAD5
M05743_2.00 |
Homo sapiens |

YGTCTAGACW |

WGTCTAGACR |
SELEX
Yin et al.(2017)
SMAD5_eDBD_HT-SELEX
|
0.966
|
0.966
|
Smad5
M11292_2.00 |
Rattus norvegicus |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SMAD5_03
|
0.974
|
0.974
|
SMAD9
M11274_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SMAD8_01
|
0.948
|
0.948
|
SMAD5
M11271_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SMAD5_02
|
0.966
|
0.966
|
Smad9
M11290_2.00 |
Rattus norvegicus |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SMAD8_02
|
0.948
|
0.948
|
Smad9
M11289_2.00 |
Mus musculus |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SMAD8_03
|
0.948
|
0.948
|
SMAD5
M11272_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SMAD5_Q5_01
|
0.966
|
0.966
|
SMAD5
M11273_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SMAD5_Q5
|
0.966
|
0.966
|
Mad
M06446_2.00 |
Drosophila melanogaster |
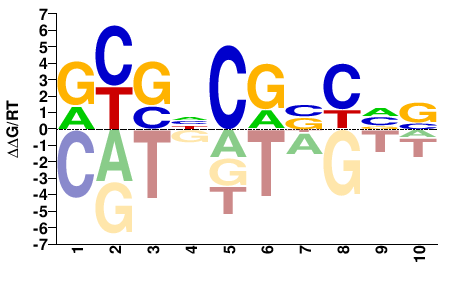

GYGHCGBCVS |

SBGVCGDCRC |
B1H
Zhu et al.(2011)
Mad_FlyReg_FBgn0011648
|
0.935
|
0.935
|
Mad
M08427_2.00 |
Drosophila melanogaster |
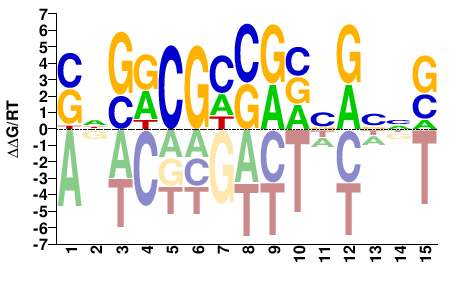

SNGRCGMSRVSRBNS |
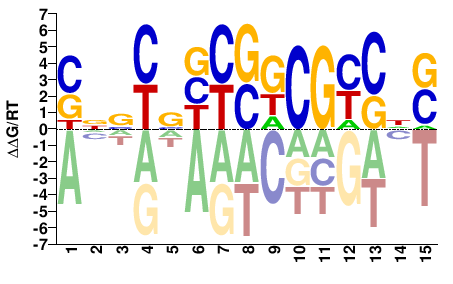
SNVYSBYSKCGYCNS |
ChIP-chip
Mathelier et al.(2014)
MA0535.1
|
0.902
|
0.902
|
SMAD5
M05744_2.00 |
Homo sapiens |

CGTCTAGACR |

YGTCTAGACG |
SELEX
Yin et al.(2017)
SMAD5_eDBD_Methyl-HT-SELEX
|
0.966
|
0.966
|
Mad
M09692_2.00 |
Drosophila melanogaster |
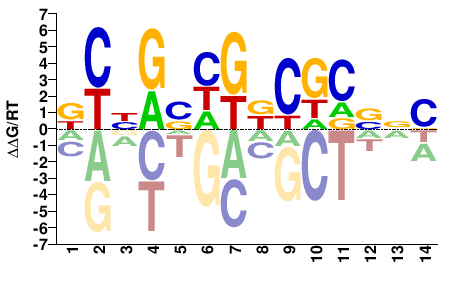
KCBGSHGKCGCSNC |
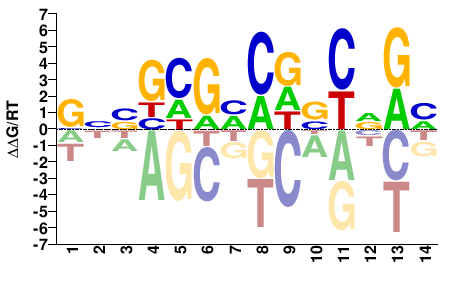

GNSGCGMCDSCVGM |
Misc
Kulakovskiy et al.(2009)
Mad
|
0.924
|
0.924
|
Mad
M11293_2.00 |
Drosophila melanogaster |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
I$MAD_Q6
|
0.904
|
0.904
|
Smad3
M00192_2.00 |
Mus musculus |
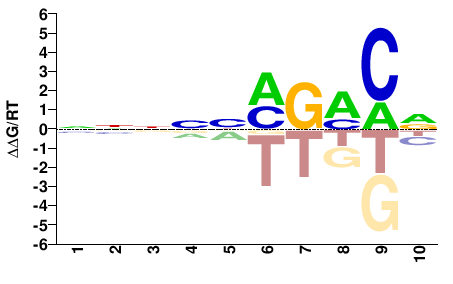
NNNNNMGMCN |
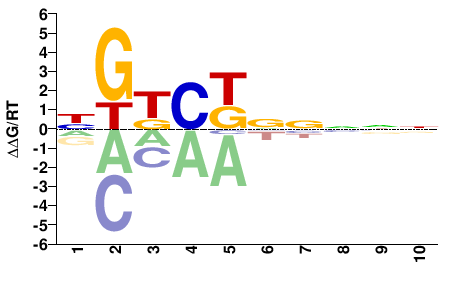
NGKCKNNNNN |
PBM
Badis et al.(2009)
Smad3_3805
|
0.701
|
0.701
|
SMAD3
M03482_2.00 |
Homo sapiens |

YGTCTAGACA |
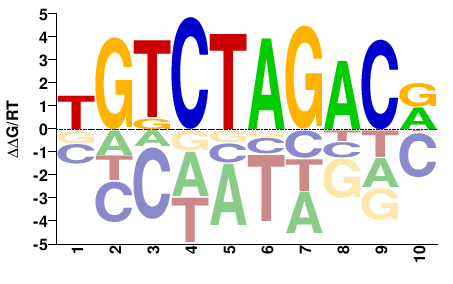

TGTCTAGACR |
SELEX
Jolma et al.(2013)
SMAD3_1
|
0.701
|
0.701
|
SMAD3
M09380_2.00 |
Homo sapiens |

NBCTSHCWSCWB |

VWGSWGDSAGVN |
Misc
Kulakovskiy et al.(2013)
SMAD3_HUMAN.H11MO.0.B
|
0.701
|
0.701
|
Smad3
M09384_2.00 |
Mus musculus |

NBCTSHCWSCWB |

VWGSWGDSAGVN |
Misc
Kulakovskiy et al.(2013)
SMAD3_MOUSE.H11MO.0.B
|
0.701
|
0.701
|
Smad3
M09637_2.00 |
Mus musculus |
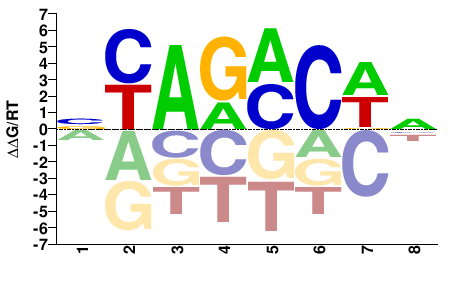
BYAGMCWN |

NWGKCTRV |
Misc
Heinz et al.(2010)
NPC-Smad3_GSE36673
|
0.701
|
0.701
|
SMAD3
M11283_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SMAD3_Q6_01
|
0.701
|
0.701
|
SMAD3
M11284_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SMAD3_Q6_02
|
0.701
|
0.701
|
SMAD3
M11285_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SMAD3_Q6
|
0.701
|
0.701
|
| For this family, TFs with SR scores > 0.700 will likely have a similar motif |