|
|
ENSP00000160740
(Macaca fascicularis)
Sox
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
| PF00505 (HMG_box) |
IPR000910 |
ENSP00000160740 |
T333700_2.00 |
GigaDB (2015-Oct-22) |
| NCBI Gene Info:The protein encoded by this gene is an ortholog of the Drosophila melanogaster capicua gene, and is a member of the high mobility group (HMG)-box superfamily of transcriptional repressors. This protein contains a conserved HMG domain that is involved in DNA binding and nuclear localization, and a conserved C-terminus. Studies suggest that the N-terminal region of this protein interacts with Atxn1 (GeneID:6310), to form a transcription repressor complex, and in vitro studies suggest that polyglutamine-expansion of ATXN1 may alter the repressor activity of this complex. Mutations in this gene have been associated with olidogdendrogliomas (PMID:21817013). In addition, translocation events resulting in gene fusions of this gene with both DUX4 (GeneID:100288687) and FOXO4 (GeneID:4303) have been associated with round cell sarcomas. There are multiple pseudogenes of this gene found on chromosomes 1, 4, 6, 7, 16, 20, and the Y chromosome. Alternative splicing results in multiple transcript variants encoding different isoforms. [provided by RefSeq, Feb 2015] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
| No direct experiments |
|
|
|
|
|
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
Cic
M00195_2.00 |
Mus musculus |
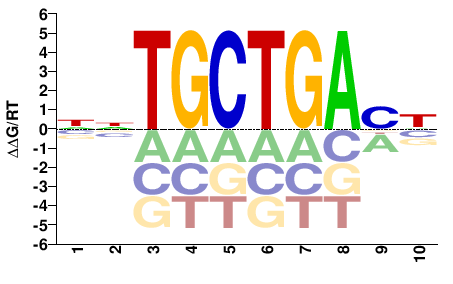
NNTGCTGABN |
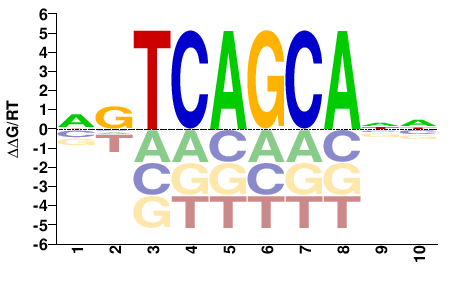

NVTCAGCANN |
PBM
Badis et al.(2009)
Cic_3454
|
0.741
|
1.000
|
cic
M06452_2.00 |
Drosophila melanogaster |

YYCATTSA |

TSAATGRR |
B1H
Zhu et al.(2011)
cic_SANGER_5_FBgn0028386
|
0.680
|
0.768
|
gei-3
M00711_2.00 |
Caenorhabditis elegans |

NWNWMWDN |
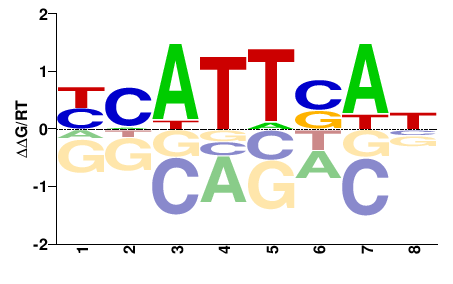

NHWKWNWN |
PBM
Narasimhan et al.(2015)
pTH10038
|
0.678
|
0.797
|
BBX
M05763_2.00 |
Homo sapiens |
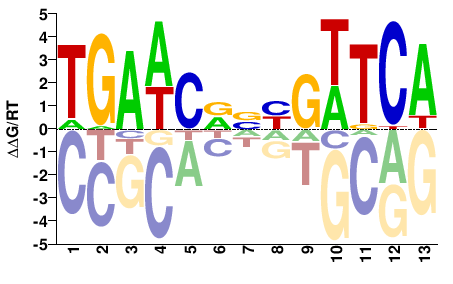
TGAWCDNYGWTCA |
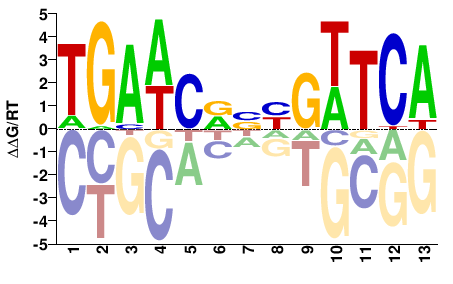
TGAWCRNHGWTCA |
SELEX
Yin et al.(2017)
BBX_eDBD_HT-SELEX
|
0.468
|
0.594
|
BBX
M05764_2.00 |
Homo sapiens |
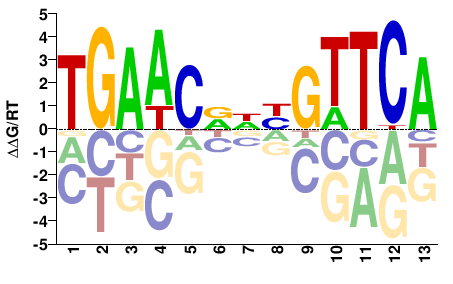
TGAACDNYGTTCA |
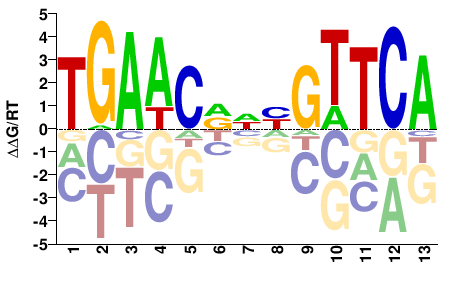
TGAACRNHGTTCA |
SELEX
Yin et al.(2017)
BBX_eDBD_Methyl-HT-SELEX
|
0.468
|
0.594
|
ROX1
M00070_2.00 |
Saccharomyces cerevisiae |
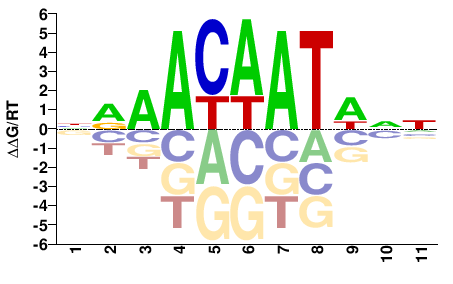
NRAACAATWNN |
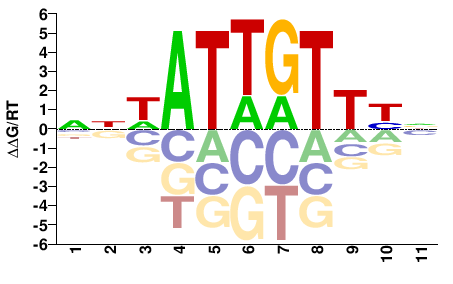
NNWATTGTTYN |
PBM
Badis et al.(2008)
ROX1_2099
|
0.440
|
0.333
|
ROX1
M07498_2.00 |
Saccharomyces cerevisiae |

YYHATTGTTYTS |

SARAACAATDRR |
PBM, CSA and or DIP-chip
Mathelier et al.(2014)
MA0371.1
|
0.440
|
0.333
|
ROX1
M08607_2.00 |
Saccharomyces cerevisiae |
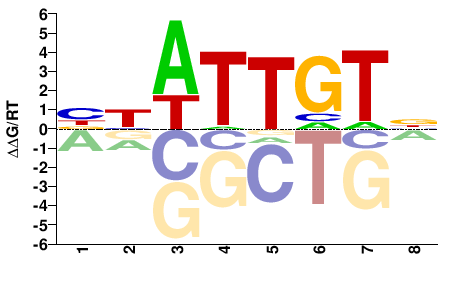
BYATTGTN |
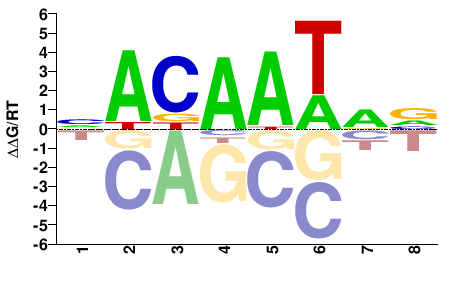
NACAATRV |
Misc
DeBoer et al.(2011)
YPR065W_1396
|
0.440
|
0.333
|
ROX1
M11346_2.00 |
Saccharomyces cerevisiae |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
F$ROX1_Q6_01
|
0.440
|
0.333
|
ROX1
M11347_2.00 |
Saccharomyces cerevisiae |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
F$ROX1_Q6
|
0.440
|
0.333
|
bbx
M03975_2.00 |
Drosophila melanogaster |
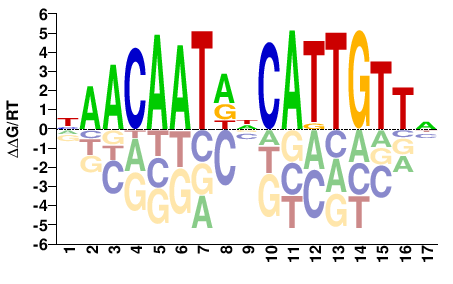
NAACAATDNCATTGTTN |

NAACAATGNHATTGTTN |
SELEX
Nitta et al.(2015)
bbx_1
|
0.440
|
0.536
|
Bbx
M00196_2.00 |
Mus musculus |

TTCATTGA |
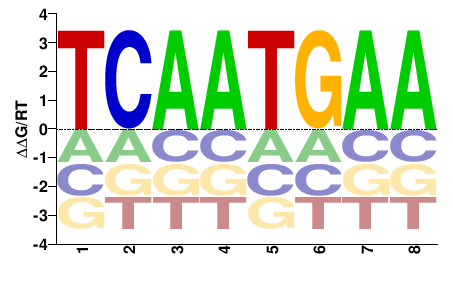

TCAATGAA |
PBM
Badis et al.(2009)
Bbx_3753
|
0.434
|
0.580
|
| For this family, TFs with SR scores > 0.415 will likely have a similar motif |