|
|
Stat3
(Mus musculus)
STAT
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
Animal TF db |
| PF02864 (STAT_bind) |
IPR013801 |
ENSMUSG00000004040 |
T337542_2.00 |
Ensembl (2018-Dec-8) |
Link out |
| NCBI Gene Info:The protein encoded by this gene is a member of the STAT protein family. In response to cytokines and growth factors, STAT family members are phosphorylated by the receptor associated kinases, and then form homo- or heterodimers that translocate to the cell nucleus where they act as transcription activators. This protein is activated through phosphorylation in response to various cytokines and growth factors including IFNs, EGF, IL5, IL6, HGF, LIF and BMP2. This protein mediates the expression of a variety of genes in response to cell stimuli, and thus plays a key role in many cellular processes such as cell growth and apoptosis. The small GTPase Rac1 has been shown to bind and regulate the activity of this protein. PIAS3 protein is a specific inhibitor of this protein. Alternative splicing results in multiple transcript variants encoding distinct isoforms. [provided by RefSeq, Sep 2015] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
Stat3
M08043_2.00 |
Mus musculus |

BTTCCYRGMAD |

HTKCYRGGAAV |
ChIP-seq
Chen et al.(2011)
SRP000217_Stat3 |
(Direct) |
(Direct) |
Stat3
M09419_2.00 |
Mus musculus |

NNYTTMCHRKMA |

TKMYDGKAARNN |
Misc
Kulakovskiy et al.(2013)
STAT3_MOUSE.H11MO.0.A |
(Direct) |
(Direct) |
Stat3
M09645_2.00 |
Mus musculus |
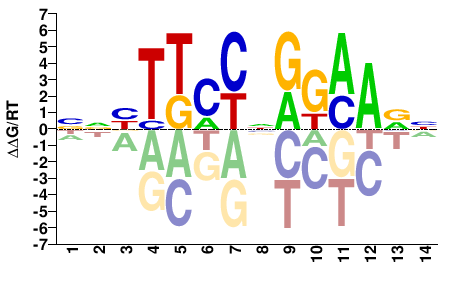
NNBTTCCNGGAAVN |
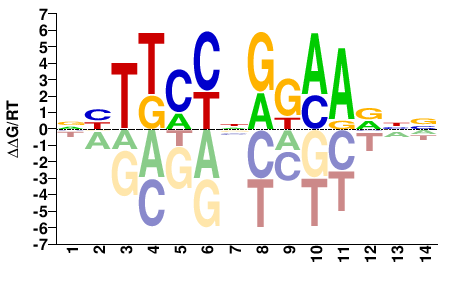
NBTTCCNGGAAVNN |
Misc
Heinz et al.(2010)
CD4-Stat3_GSE19198 |
(Direct) |
(Direct) |
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
STAT3
M08172_2.00 |
Homo sapiens |
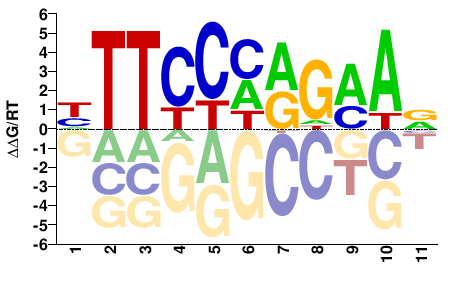
HTTCCHRGMAV |

BTKCYDGGAAD |
ChIP-seq
Mathelier et al.(2014)
MA0144.2
|
1.000
|
1.000
|
STAT3
M08015_2.00 |
Homo sapiens |
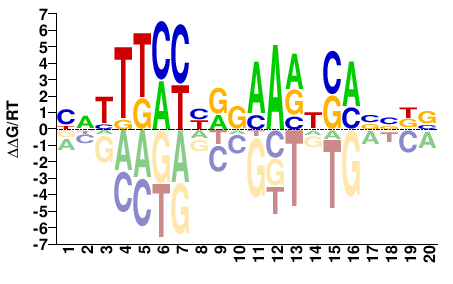
YDTTTMYYRGAARHSABVDB |
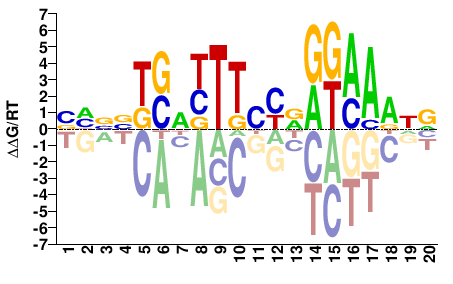
VHBVTSDYTTCYRRKAAAHR |
ChIP-seq
Gerstein et al.(2012)
GM12878_STAT3_Stanford
|
1.000
|
1.000
|
STAT3
M08016_2.00 |
Homo sapiens |
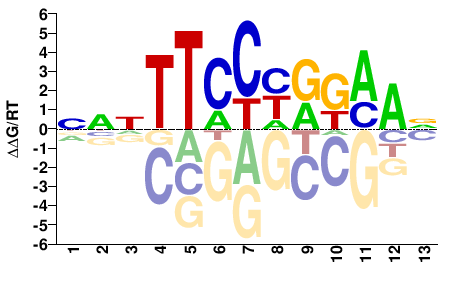
NNNTTCCHRKAAN |

NTTMYDGGAANNN |
ChIP-seq
Gerstein et al.(2012)
HeLa-S3_STAT3_Stanford
|
1.000
|
1.000
|
STAT3
M08017_2.00 |
Homo sapiens |
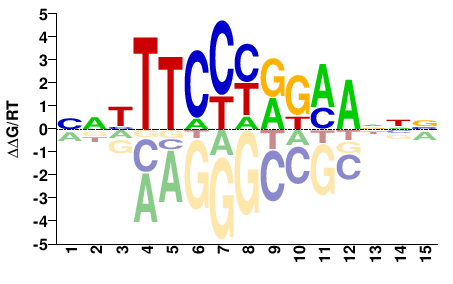
NNHTTCCYRKMANNN |
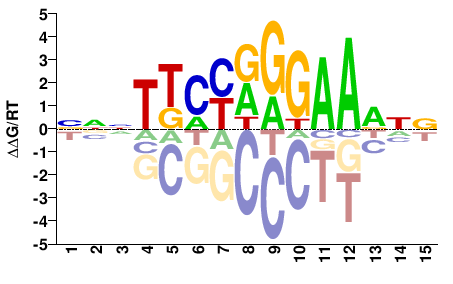
NNNTKMYRGGAADNN |
ChIP-seq
Gerstein et al.(2012)
MCF10A-Er-Src_STAT3_Harvard#Weissman
|
1.000
|
1.000
|
STAT3
M08018_2.00 |
Homo sapiens |
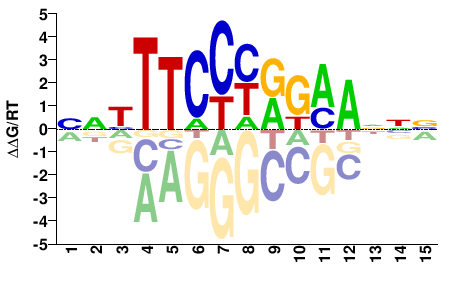
NNHTTCCYRKMANNN |
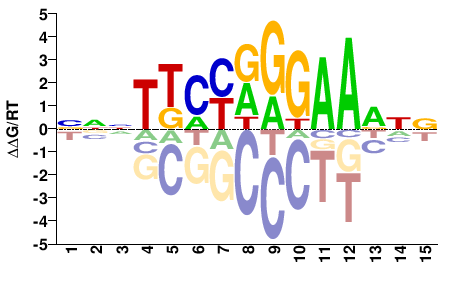
NNNTKMYRGGAADNN |
ChIP-seq
Gerstein et al.(2012)
MCF10A-Er-Src_STAT3_Stanford
|
1.000
|
1.000
|
STAT3
M09415_2.00 |
Homo sapiens |
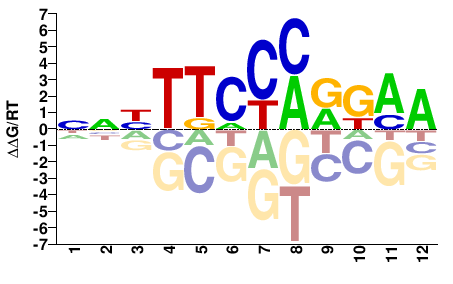
NNYTTCCMRGAA |

TTCYKGGAARNN |
Misc
Kulakovskiy et al.(2013)
STAT3_HUMAN.H11MO.0.A
|
1.000
|
1.000
|
STAT3
M11367_2.00 |
Homo sapiens |
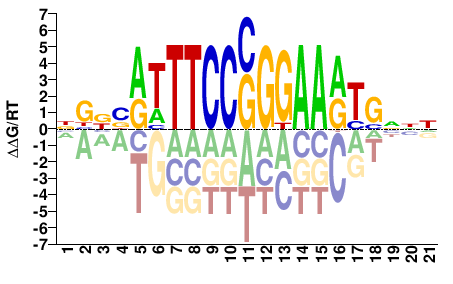
NBBBATTTCCSGGAARTGNNN |
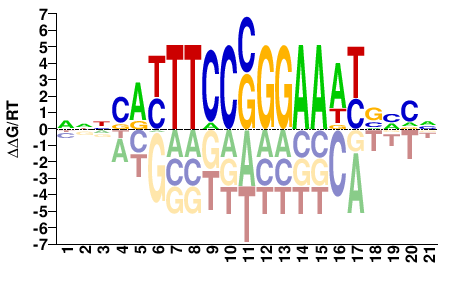
NNNCAYTTCCSGGAAATVVVN |
Transfac
Matys et al.(2006)
V$STAT3_01
|
1.000
|
1.000
|
STAT3
M11368_2.00 |
Homo sapiens |
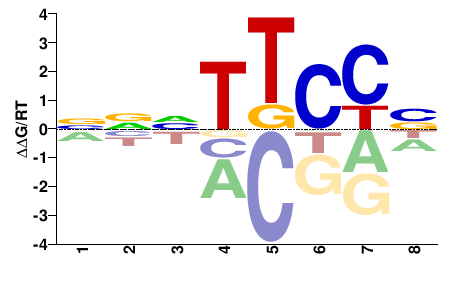
NNNTTCCN |
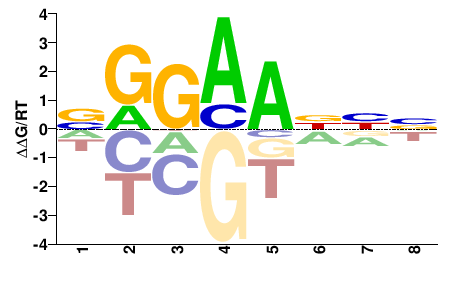
NGGAANNN |
Transfac
Matys et al.(2006)
V$STAT3_02
|
1.000
|
1.000
|
STAT3
M11369_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$STAT3_03
|
1.000
|
1.000
|
STAT3
M11370_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$STAT3_Q4
|
1.000
|
1.000
|
| For this family, TFs with SR scores > 0.700 will likely have a similar motif |