|
|
TBP
(Homo sapiens)
TBP
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
Animal TF db |
| PF00352 (TBP) |
IPR000814 |
ENSG00000112592 |
T342320_2.00 |
Ensembl (2018-Dec-8) |
Link out |
| NCBI Gene Info:Initiation of transcription by RNA polymerase II requires the activities of more than 70 polypeptides. The protein that coordinates these activities is transcription factor IID (TFIID), which binds to the core promoter to position the polymerase properly, serves as the scaffold for assembly of the remainder of the transcription complex, and acts as a channel for regulatory signals. TFIID is composed of the TATA-binding protein (TBP) and a group of evolutionarily conserved proteins known as TBP-associated factors or TAFs. TAFs may participate in basal transcription, serve as coactivators, function in promoter recognition or modify general transcription factors (GTFs) to facilitate complex assembly and transcription initiation. This gene encodes TBP, the TATA-binding protein. A distinctive feature of TBP is a long string of glutamines in the N-terminus. This region of the protein modulates the DNA binding activity of the C terminus, and modulation of DNA binding affects the rate of transcription complex formation and initiation of transcription. The number of CAG repeats encoding the polyglutamine tract is usually 25-42, and expansion of the number of repeats to 45-66 increases the length of the polyglutamine string and is associated with spinocerebellar ataxia 17, a neurodegenerative disorder classified as a polyglutamine disease. Two transcript variants encoding different isoforms have been found for this gene. [provided by RefSeq, Jul 2016] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
TBP
M08231_2.00 |
Homo sapiens |

YATTTGCATD |

HATGCAAATR |
ChIP-seq
Contrino et al.(2012)
Mv129 |
(Direct) |
(Direct) |
TBP
M09432_2.00 |
Homo sapiens |
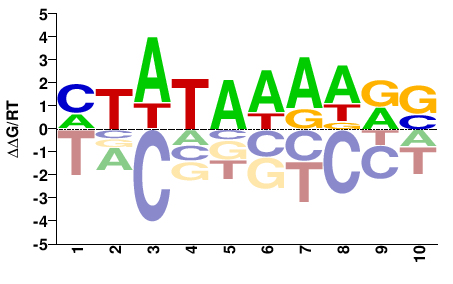

MTATAWAWRS |

SYWTWTATAK |
Misc
Kulakovskiy et al.(2013)
TBP_HUMAN.H11MO.0.A |
(Direct) |
(Direct) |
TBP
M11388_2.00 |
Homo sapiens |
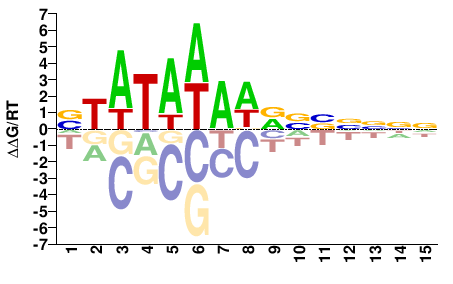
VTATAWAWRVVNNNN |
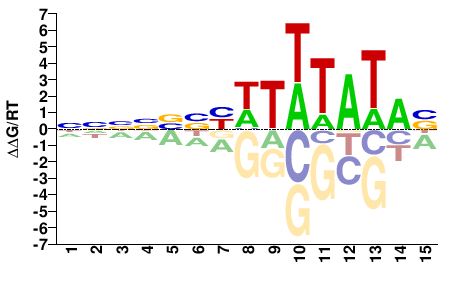
NNNNBBYWTWTATAB |
Transfac
Matys et al.(2006)
V$TATA_01 |
(Direct) |
(Direct) |
TBP
M11389_2.00 |
Homo sapiens |

NCYWTAAAAD |

HTTTTAWRGN |
Transfac
Matys et al.(2006)
V$TATA_C |
(Direct) |
(Direct) |
TBP
M11390_2.00 |
Homo sapiens |

KATAAATW |
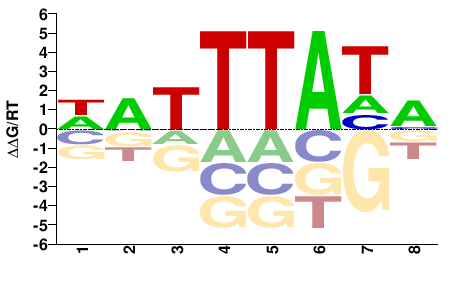
WATTTATM |
Transfac
Matys et al.(2006)
V$TBP_01 |
(Direct) |
(Direct) |
TBP
M11391_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$TBP_Q6_01 |
(Direct) |
(Direct) |
TBP
M11392_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$TBP_Q6 |
(Direct) |
(Direct) |
TBP
M11491_2.00 |
Homo sapiens |
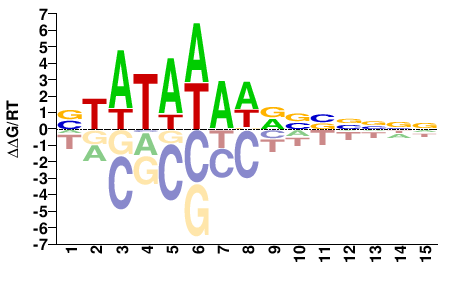
VTATAWAWRVVNNNN |
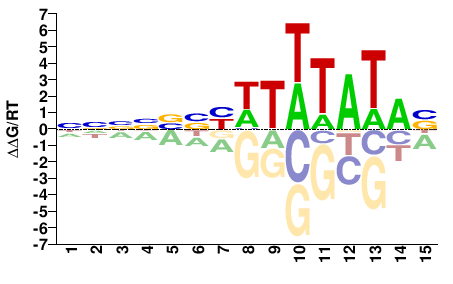
NNNNBBYWTWTATAB |
Unknown
Mathelier et al.(2014)
MA0108.2 |
(Direct) |
(Direct) |
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
Tbp
M00216_2.00 |
Mus musculus |
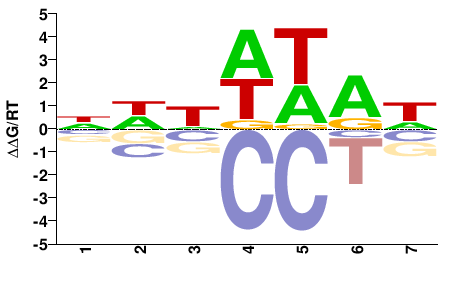
NWWWWAW |
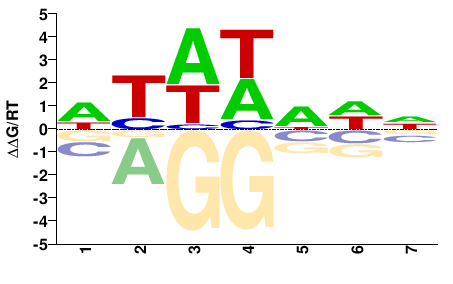
WTWWWWN |
PBM
Badis et al.(2009)
Tbp_pr781
|
1.000
|
1.000
|
Tbp
M09433_2.00 |
Mus musculus |

VTATAAAARS |

SYTTTTATAB |
Misc
Kulakovskiy et al.(2013)
TBP_MOUSE.H11MO.0.A
|
1.000
|
1.000
|
SPT15
M01567_2.00 |
Saccharomyces cerevisiae |
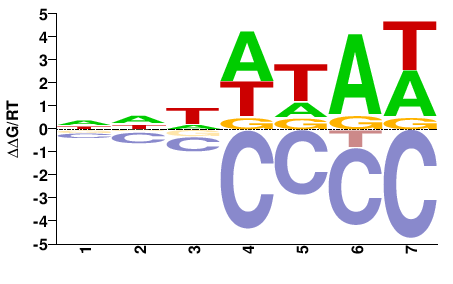
NNDWDAW |
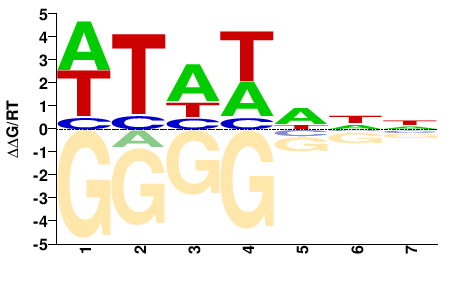
WTHWHNN |
PBM
Zhu et al.(2009)
Spt15
|
0.808
|
0.808
|
SPT15
M08609_2.00 |
Saccharomyces cerevisiae |
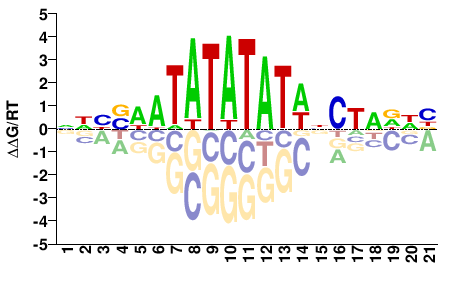
NNBBHWTATATATWNCNNDNB |
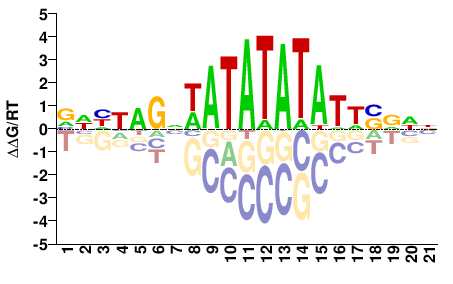
VNHNNGNWATATATAWDVVNN |
Misc
DeBoer et al.(2011)
YER148W_798
|
0.808
|
0.808
|
SPT15
M11393_2.00 |
Saccharomyces cerevisiae |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
F$TBP_Q6
|
0.808
|
0.808
|
| For this family, TFs with SR scores > 0.700 will likely have a similar motif |