|
|
HHAL003746-PA
(Halyomorpha halys)
TCR/CxC
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
| PF03638 (TCR) |
IPR005172 |
HHAL003746-PA |
T346179_2.00 |
Misc (2018-Jan-19) |
| NCBI Gene Info:This gene encodes a tumor suppressor protein containing transcriptional activation, DNA binding, and oligomerization domains. The encoded protein responds to diverse cellular stresses to regulate expression of target genes, thereby inducing cell cycle arrest, apoptosis, senescence, DNA repair, or changes in metabolism. Mutations in this gene are associated with a variety of human cancers, including hereditary cancers such as Li-Fraumeni syndrome. Alternative splicing of this gene and the use of alternate promoters result in multiple transcript variants and isoforms. Additional isoforms have also been shown to result from the use of alternate translation initiation codons from identical transcript variants (PMIDs: 12032546, 20937277). [provided by RefSeq, Dec 2016] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
| No direct experiments |
|
|
|
|
|
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
LIN54
M02527_2.00 |
Gallus gallus |

NRTTYAAAH |

DTTTRAAYN |
PBM
Weirauch et al.(2014)
pTH8566
|
0.672
|
0.797
|
SPU_025306
M02531_2.00 |
Strongylocentrotus purpuratus |
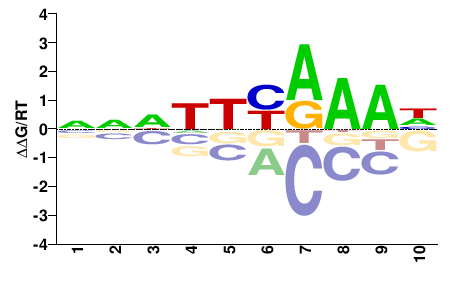
NNNNWYRAAH |
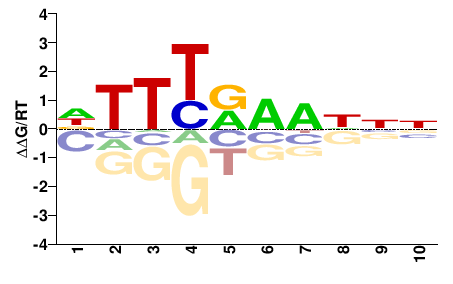
DTTYRWNNNN |
PBM
Weirauch et al.(2014)
pTH9286
|
0.672
|
0.797
|
GE12548
M02529_2.00 |
Drosophila yakuba |

DTTYAAAY |

RTTTRAAH |
PBM
Weirauch et al.(2014)
pTH9262
|
0.672
|
0.781
|
BGIBMGA008956
M02524_2.00 |
Bombyx mori |

AATTTAAAW |

WTTTAAATT |
PBM
Weirauch et al.(2014)
pTH9238
|
0.672
|
0.766
|
lin54
M02526_2.00 |
Danio rerio |
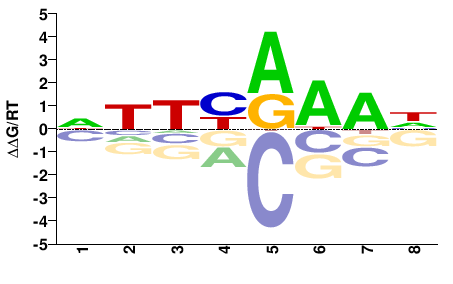
NHTYRAAH |
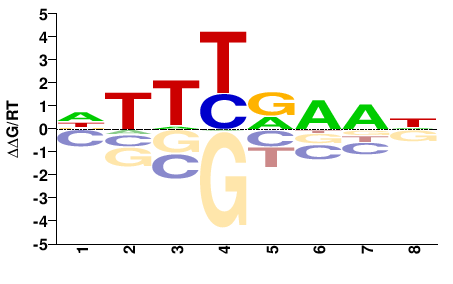
DTTYRADN |
PBM
Weirauch et al.(2014)
pTH8399
|
0.672
|
0.766
|
ENSTSYG00000005291
M02528_2.00 |
Tarsius syrichta |

NNWDHDHWNN |

NNWDHDHWNN |
PBM
Weirauch et al.(2014)
pTH9366
|
0.672
|
0.766
|
NEMVEDRAFT_v1g111410
M01320_2.00 |
Nematostella vectensis |
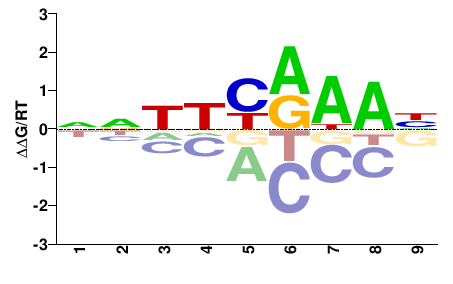
NNNNYRWRN |
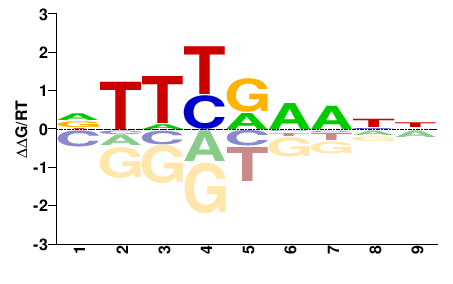
NYWYRNNNN |
PBM
Lambert et al.(2019)
pTH9771
|
0.672
|
0.719
|
XP_002124866.1
M01321_2.00 |
Ciona intestinalis |

NDTTYRAAN |

NTTYRAAHN |
PBM
Lambert et al.(2019)
pTH9835
|
0.672
|
0.719
|
AT2G20110
M02520_2.00 |
Arabidopsis thaliana |

DTTTAAAH |

DTTTAAAH |
PBM
Weirauch et al.(2014)
pTH7914
|
0.672
|
0.625
|
TCX5
M02523_2.00 |
Arabidopsis thaliana |

NRTTCAAAYN |

NRTTTGAAYN |
PBM
Weirauch et al.(2014)
pTH8164
|
0.672
|
0.625
|
fgeneshCV_pg.C_scaffold_18000164
M02534_2.00 |
Chlorella vulgaris |
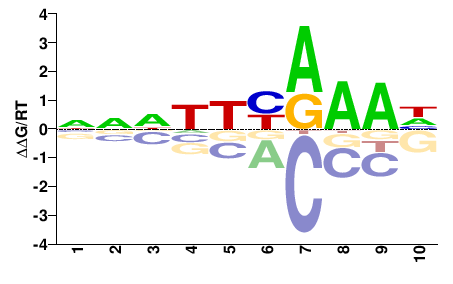
NNNNDYRAAH |
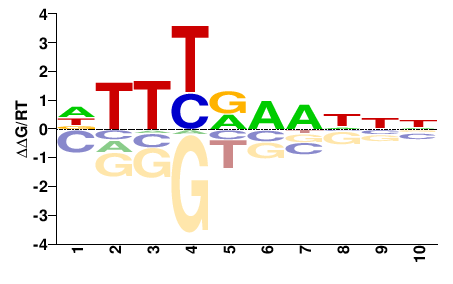
DTTYRHNNNN |
PBM
Weirauch et al.(2014)
pTH9252
|
0.672
|
0.625
|
AT2G20110
M07295_2.00 |
Arabidopsis thaliana |
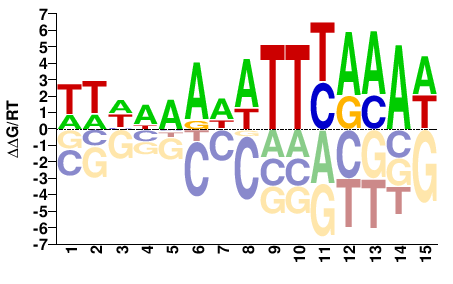
WWHWAADATTYAAAW |

WTTTRAATHTTWDWW |
Dap-seq
OMalley et al.(2016)
AT2G20110_col_a
|
0.672
|
0.625
|
AT2G20110
M07296_2.00 |
Arabidopsis thaliana |

WWHNAAATTYAAAHN |

NDTTTRAATTTNDWW |
Dap-seq
OMalley et al.(2016)
AT2G20110_colamp_a
|
0.672
|
0.625
|
AT5G25790
M01319_2.00 |
Arabidopsis thaliana |
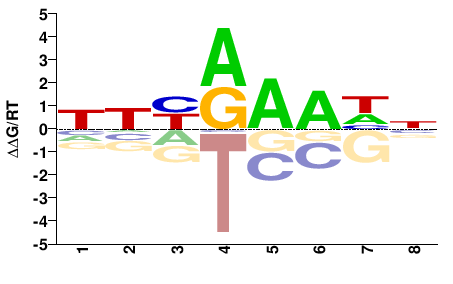
NHYRAAHN |
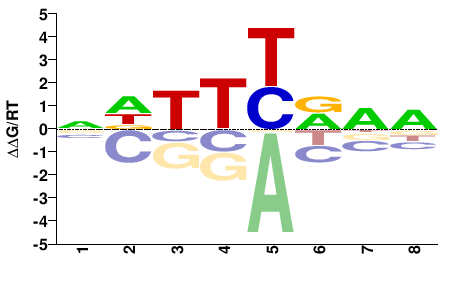

NDTTYRDN |
PBM
Lambert et al.(2019)
pTH11533
|
0.672
|
0.609
|
TCX4
M02521_2.00 |
Arabidopsis thaliana |

DWTTYRAAWH |

DWTTYRAAWH |
PBM
Weirauch et al.(2014)
pTH7573
|
0.672
|
0.547
|
TCX3
M01087_2.00 |
Arabidopsis thaliana |
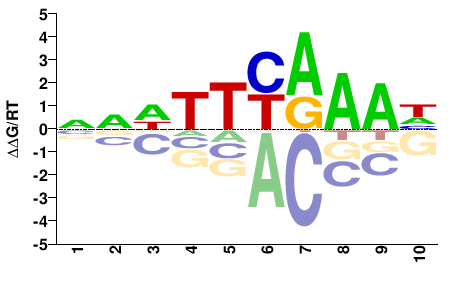
NNDTTYAAAH |
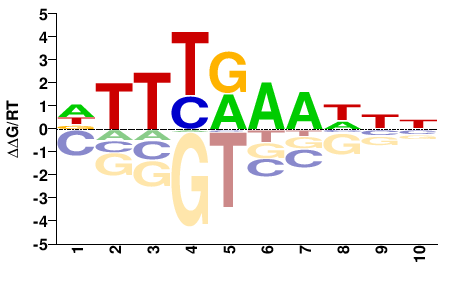
DTTTRAAHNN |
PBM
Sullivan et al.(2014)
pTH7218
|
0.672
|
0.531
|
TCX3
M02522_2.00 |
Arabidopsis thaliana |

WTTYAAAH |
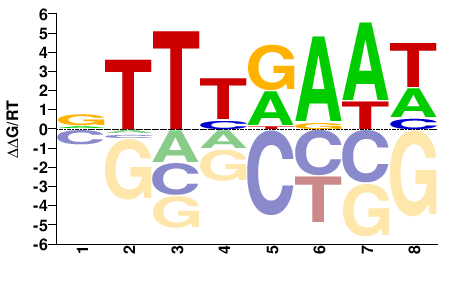
DTTTRAAW |
PBM
Weirauch et al.(2014)
pTH7421
|
0.672
|
0.531
|
TCX3
M07297_2.00 |
Arabidopsis thaliana |
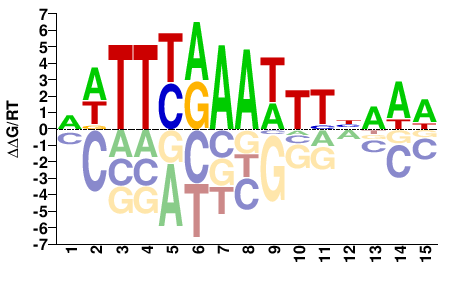
DWTTYRAAWTTBAAW |
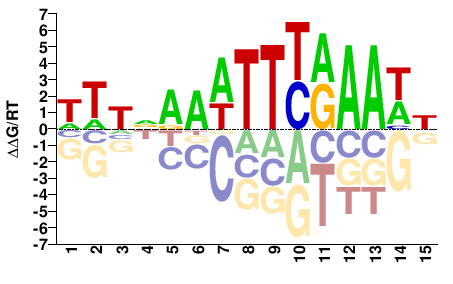
WTTVAAATTYRAAWH |
Dap-seq
OMalley et al.(2016)
SOL1_colamp_a
|
0.672
|
0.531
|
TCX3
M07298_2.00 |
Arabidopsis thaliana |
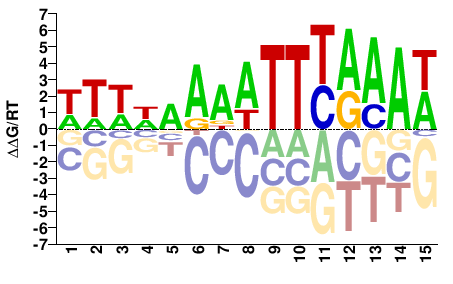
WTWWAAAATTYAAAW |

WTTTRAATTTTWWAW |
Dap-seq
OMalley et al.(2016)
SOL1_col
|
0.672
|
0.531
|
TCX2
M07300_2.00 |
Arabidopsis thaliana |
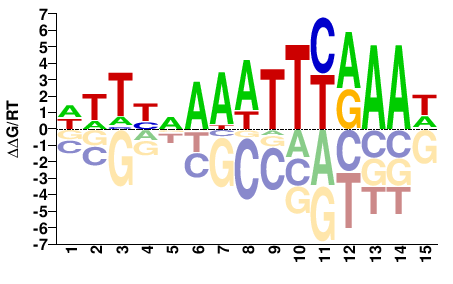
WWTYVAAATTYRAAW |

WTTYRAATTTBRAWW |
Dap-seq
OMalley et al.(2016)
TCX2_col_a
|
0.672
|
0.531
|
TCX2
M07301_2.00 |
Arabidopsis thaliana |

TTYVAAATTYRAAW |

WTTYRAATTTBRAA |
Dap-seq
OMalley et al.(2016)
TCX2_colamp_a
|
0.672
|
0.531
|
BRADI1G55710
M02525_2.00 |
Brachypodium distachyon |

NVTBYRVABN |

NVTBYRVABN |
PBM
Weirauch et al.(2014)
pTH9241
|
0.672
|
0.516
|
PK22848.1
M02532_2.00 |
Cannabis sativa |
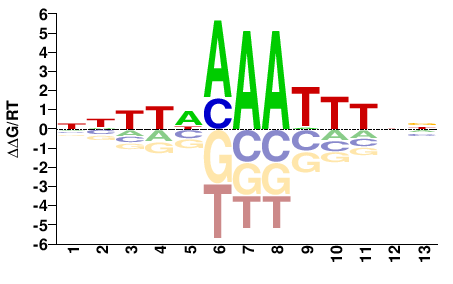
NNNYNAAATTTNN |
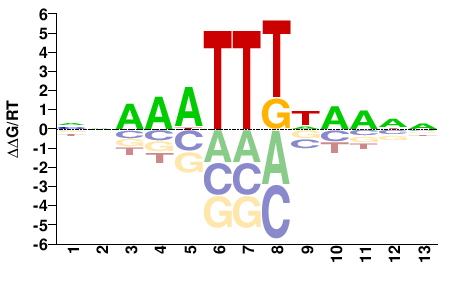
NNAAATTTNRNNN |
PBM
Weirauch et al.(2014)
pTH9724
|
0.672
|
0.516
|
fgenesh1_pg.C_Chr_09.0001000301
M02533_2.00 |
Ostreococcus tauri |
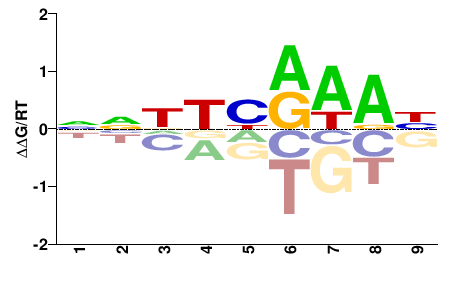
NNNNNRHNN |
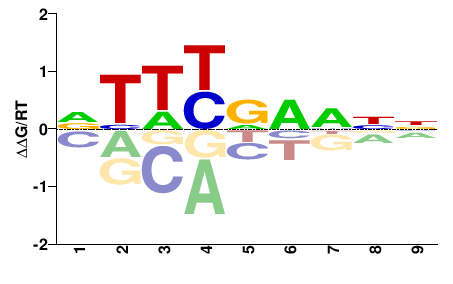
NNDYNNNNN |
PBM
Weirauch et al.(2014)
pTH9457
|
0.672
|
0.516
|
gw1.20.00.89.1
M02535_2.00 |
Ostreococcus tauri |

WNYWBVWRNW |

WNYWBVWRNW |
PBM
Weirauch et al.(2014)
pTH9647
|
0.672
|
0.516
|
GSPATT00010696001
M02530_2.00 |
Paramecium tetraurelia |
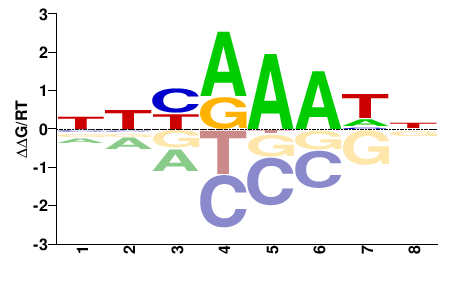
NNBRAAHN |
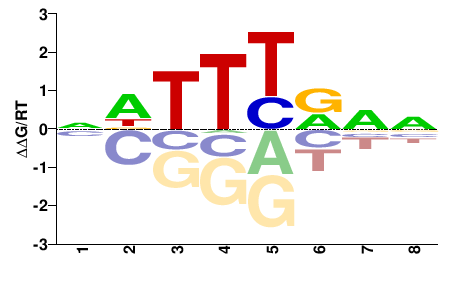
NDTTYVNN |
PBM
Weirauch et al.(2014)
pTH8720
|
0.672
|
0.500
|
PK27683.1
M01322_2.00 |
Cannabis sativa |
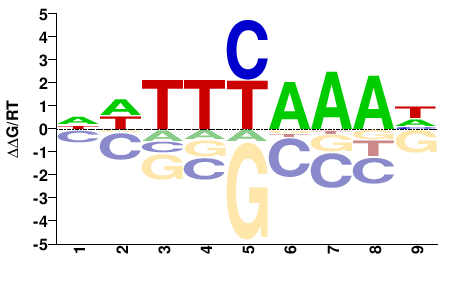
NDTTYAAAH |

DTTTRAAHN |
PBM
Lambert et al.(2019)
pTH11347
|
0.672
|
0.500
|
TSO1
M07299_2.00 |
Arabidopsis thaliana |
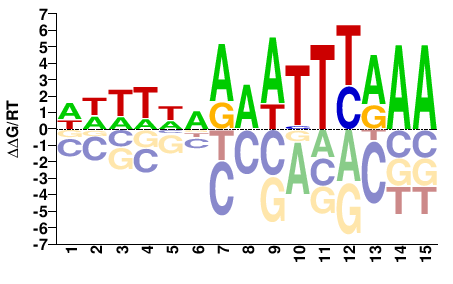
WWWTWDAAATTYAAA |

TTTRAATTTHWAWWW |
Dap-seq
OMalley et al.(2016)
TSO1_col_a
|
0.672
|
0.500
|
| For this family, TFs with SR scores > 0.660 will likely have a similar motif |