|
|
FOS
(Homo sapiens)
bZIP
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
Animal TF db |
| PF00170 (bZIP_1) |
IPR011616 |
ENSG00000170345 |
T059755_2.00 |
Ensembl (2018-Dec-8) |
Link out |
| NCBI Gene Info:The Fos gene family consists of 4 members: FOS, FOSB, FOSL1, and FOSL2. These genes encode leucine zipper proteins that can dimerize with proteins of the JUN family, thereby forming the transcription factor complex AP-1. As such, the FOS proteins have been implicated as regulators of cell proliferation, differentiation, and transformation. In some cases, expression of the FOS gene has also been associated with apoptotic cell death. [provided by RefSeq, Jul 2008] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
FOS
M04327_2.00 |
Homo sapiens |

BRTGACGTCAYV |

BRTGACGTCAYV |
SELEX
Yin et al.(2017)
FOS_eDBD_HT-SELEX |
(Direct) |
(Direct) |
FOS
M04328_2.00 |
Homo sapiens |

VATGACRTCAYM |

KRTGAYGTCATB |
SELEX
Yin et al.(2017)
FOS_eDBD_Methyl-HT-SELEX |
(Direct) |
(Direct) |
FOS
M07826_2.00 |
Homo sapiens |

NATGASTCABDBN |
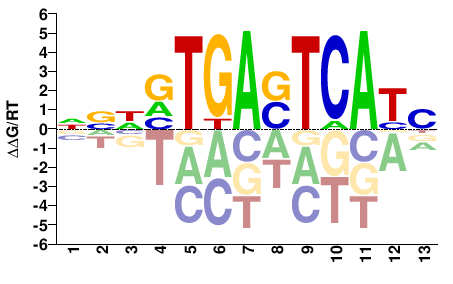
NVHVTGASTCATN |
ChIP-seq
Gerstein et al.(2012)
HeLa-S3_CFOS_Stanford |
(Direct) |
(Direct) |
FOS
M07827_2.00 |
Homo sapiens |

MTGASTCAYHN |

NDRTGASTCAK |
ChIP-seq
Gerstein et al.(2012)
HUVEC_CFOS_UCD |
(Direct) |
(Direct) |
FOS
M07828_2.00 |
Homo sapiens |

BDRTGASTCAYNBHB |
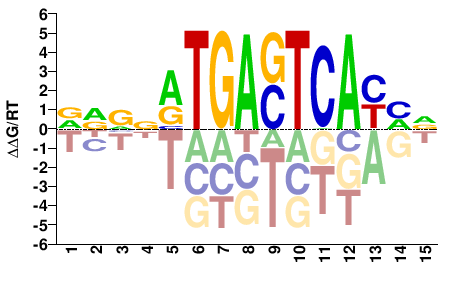
VDVNRTGASTCAYHV |
ChIP-seq
Gerstein et al.(2012)
K562_CFOS_UChicago |
(Direct) |
(Direct) |
FOS
M08069_2.00 |
Homo sapiens |

DVTGASTCATN |

NATGASTCABH |
ChIP-seq
Mathelier et al.(2014)
MA0476.1 |
(Direct) |
(Direct) |
FOS
M08802_2.00 |
Homo sapiens |
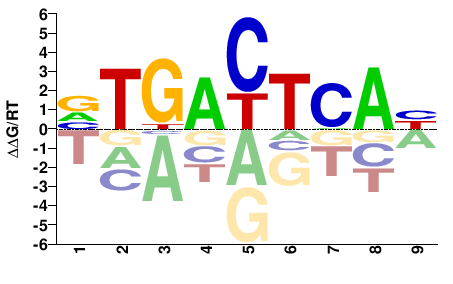

VTGACTCAB |

VTGAGTCAB |
Misc
Kulakovskiy et al.(2013)
FOS_HUMAN.H11MO.0.A |
(Direct) |
(Direct) |
FOS
M09988_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$CFOS_Q4 |
(Direct) |
(Direct) |
FOS
M09989_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$CFOS_Q6 |
(Direct) |
(Direct) |
FOS
M09990_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$FOS_02 |
(Direct) |
(Direct) |
FOS
M09991_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$FOS_03 |
(Direct) |
(Direct) |
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
Fos
M08819_2.00 |
Mus musculus |

NRTGACTCAYNN |
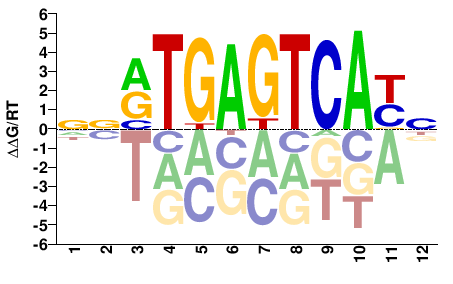
NNRTGAGTCAYN |
Misc
Kulakovskiy et al.(2013)
FOS_MOUSE.H11MO.0.A
|
0.944
|
1.000
|
Fosl2
M01809_2.00 |
Mus musculus |
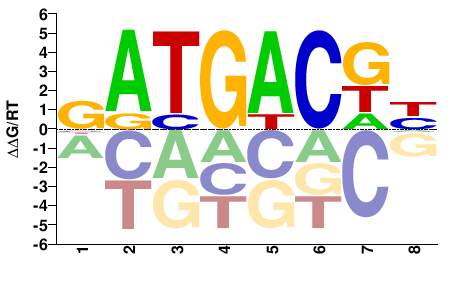
GATGACKH |

DMGTCATC |
PBM
Weirauch et al.(2014)
pTH5108
|
0.891
|
0.810
|
Fosl2
M08828_2.00 |
Mus musculus |
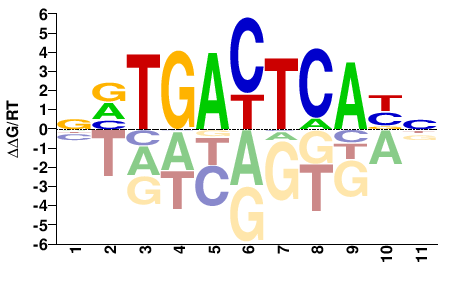

NVTGACTCABN |

NVTGAGTCABN |
Misc
Kulakovskiy et al.(2013)
FOSL2_MOUSE.H11MO.0.A
|
0.891
|
0.810
|
FOSB
M04272_2.00 |
Homo sapiens |

NRTGACGTCAYN |

NRTGACGTCAYN |
SELEX
Yin et al.(2017)
FOSB_eDBD_HT-SELEX
|
0.810
|
0.909
|
Fosl2
M09494_2.00 |
Mus musculus |
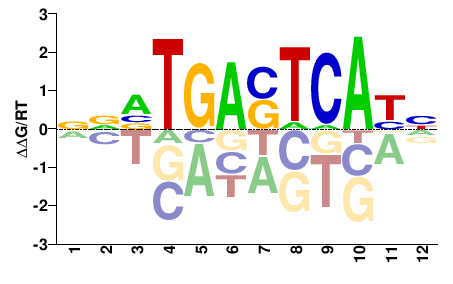

NNVTGASTCABN |

NVTGASTCABNN |
Misc
Heinz et al.(2010)
3T3L1-Fosl2_GSE56872
|
0.891
|
0.810
|
FOSB
M08795_2.00 |
Homo sapiens |
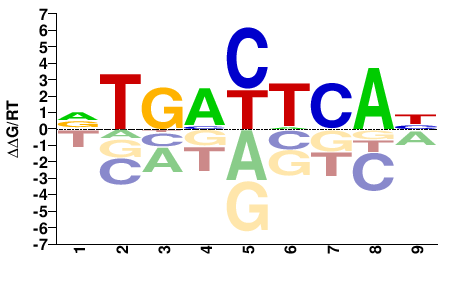

VTGACTCAB |

VTGAGTCAB |
Misc
Kulakovskiy et al.(2013)
FOSB_HUMAN.H11MO.0.A
|
0.810
|
0.909
|
Fosb
M08814_2.00 |
Mus musculus |
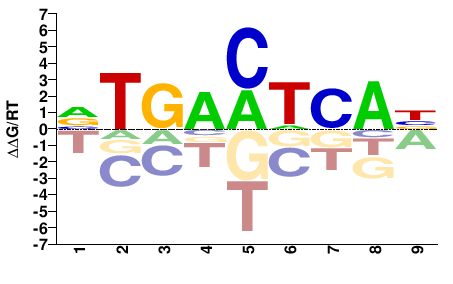

VTGACTCAB |

VTGAGTCAB |
Misc
Kulakovskiy et al.(2013)
FOSB_MOUSE.H11MO.0.A
|
0.810
|
0.909
|
FOSB
M04273_2.00 |
Homo sapiens |

NRTGACRTCAYN |

NRTGAYGTCAYN |
SELEX
Yin et al.(2017)
FOSB_eDBD_Methyl-HT-SELEX
|
0.810
|
0.882
|
Fosl1
M01805_2.00 |
Mus musculus |
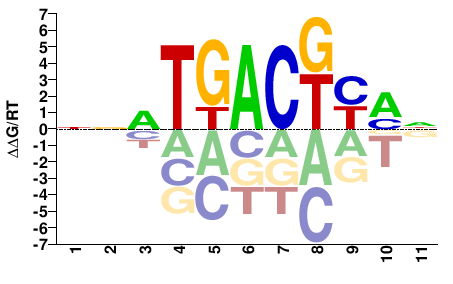
NNRTGACKYMN |
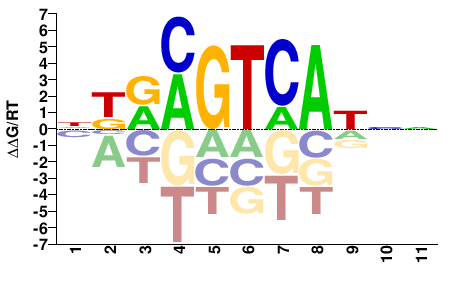

NKRMGTCAYNN |
PBM
Weirauch et al.(2014)
pTH5077
|
0.823
|
0.762
|
FOSL1
M04333_2.00 |
Homo sapiens |
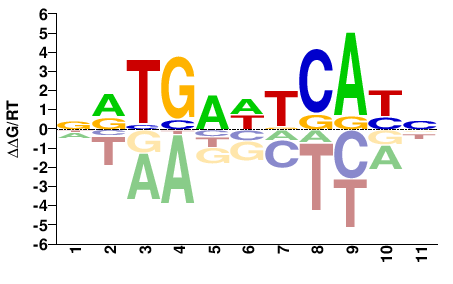
NRTGAWTCAYN |
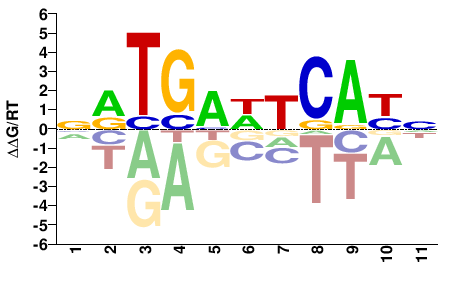

NRTGAWTCAYN |
SELEX
Yin et al.(2017)
FOSL1_FL_HT-SELEX_1
|
0.823
|
0.762
|
FOSL1
M04334_2.00 |
Homo sapiens |

NRTGACGTCAYN |

NRTGACGTCAYN |
SELEX
Yin et al.(2017)
FOSL1_FL_HT-SELEX_2
|
0.823
|
0.762
|
FOSL1
M04335_2.00 |
Homo sapiens |

DRTGAYRCR |

YGYRTCAYH |
SELEX
Yin et al.(2017)
FOSL1_FL_HT-SELEX_3
|
0.823
|
0.762
|
FOSL1
M04035_2.00 |
Homo sapiens |

RTGACGTCAY |

RTGACGTCAY |
SELEX
Rodriguez-Martinez et al.(2017)
FOSL1.1
|
0.823
|
0.762
|
FOSL1
M04036_2.00 |
Homo sapiens |
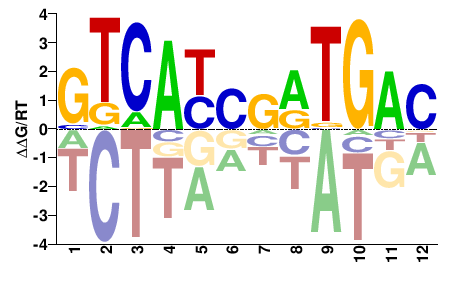
GTCAYCRRTGAC |

GTCAYYGRTGAC |
SELEX
Rodriguez-Martinez et al.(2017)
FOSL1.2
|
0.823
|
0.762
|
FOSL1
M08071_2.00 |
Homo sapiens |

DRTGASTCAKV |

BMTGASTCAYH |
ChIP-seq
Mathelier et al.(2014)
MA0477.1
|
0.823
|
0.762
|
FOSL1
M07833_2.00 |
Homo sapiens |

NDRTGASTCAB |

VTGASTCAYHN |
ChIP-seq
Gerstein et al.(2012)
K562_FOSL1_HudsonAlpha
|
0.823
|
0.762
|
FOSL1
M08805_2.00 |
Homo sapiens |
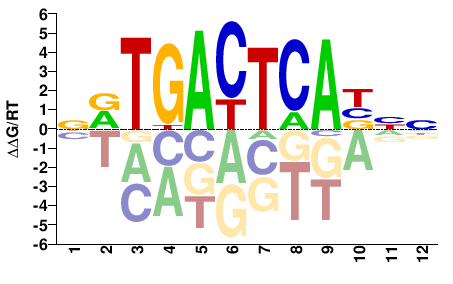
NVTGACTCABNN |

NNVTGAGTCABN |
Misc
Kulakovskiy et al.(2013)
FOSL1_HUMAN.H11MO.0.A
|
0.823
|
0.762
|
Fosl1
M08821_2.00 |
Mus musculus |
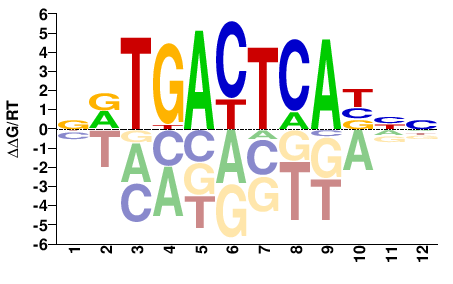
NVTGACTCABNN |

NNVTGAGTCABN |
Misc
Kulakovskiy et al.(2013)
FOSL1_MOUSE.H11MO.0.A
|
0.823
|
0.762
|
FOSL1
M09490_2.00 |
Homo sapiens |
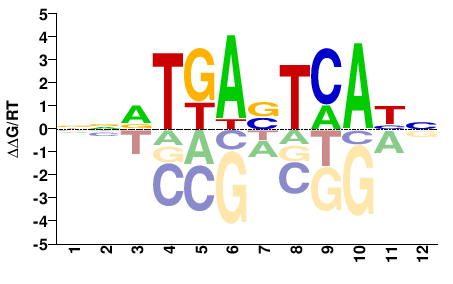
NNVTGASTCABN |

NVTGASTCABNN |
Misc
Heinz et al.(2010)
BT549-Fra1_GSE46166
|
0.823
|
0.762
|
FOSL1
M10000_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$FOSL1_01
|
0.823
|
0.762
|
FOSL1
M10001_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$FRA1_Q5
|
0.823
|
0.762
|
FOSL1
M10002_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$FRA1_Q6_01
|
0.823
|
0.762
|
FOSL1
M10003_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$FRA1_Q6
|
0.823
|
0.762
|
FOSL1
M04336_2.00 |
Homo sapiens |

BRTGASTCATV |

BATGASTCAYV |
SELEX
Yin et al.(2017)
FOSL1_FL_Methyl-HT-SELEX_1
|
0.823
|
0.762
|
FOSL1
M04337_2.00 |
Homo sapiens |

NRTSACGTCAYN |

NRTGACGTSAYN |
SELEX
Yin et al.(2017)
FOSL1_FL_Methyl-HT-SELEX_2
|
0.823
|
0.762
|
FOSL1
M04338_2.00 |
Homo sapiens |

RATGAYRCR |

YGYRTCATY |
SELEX
Yin et al.(2017)
FOSL1_FL_Methyl-HT-SELEX_3
|
0.823
|
0.762
|
| For this family, TFs with SR scores > 0.782 will likely have a similar motif |