|
|
Rela
(Mus musculus)
Rel
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
Animal TF db |
| PF00554 (RHD_DNA_bind) |
IPR011539 |
ENSMUSG00000024927 |
T317313_2.00 |
Ensembl (2018-Dec-8) |
Link out |
| NCBI Gene Info:Enables several functions, including DNA-binding transcription factor activity, RNA polymerase II-specific; actinin binding activity; and ankyrin repeat binding activity. Involved in several processes, including negative regulation of protein sumoylation; postsynapse to nucleus signaling pathway; and regulation of transcription, DNA-templated. Acts upstream of or within several processes, including cellular response to hepatocyte growth factor stimulus; cellular response to lipopolysaccharide; and positive regulation of interleukin-12 production. Located in several cellular components, including chromatin; cytosol; and glutamatergic synapse. Is expressed in several structures, including alimentary system; early conceptus; genitourinary system; musculature; and skin. Human ortholog(s) of this gene implicated in ductal carcinoma in situ; lung non-small cell carcinoma; lymphoma; and renal cell carcinoma. Orthologous to human RELA (RELA proto-oncogene, NF-kB subunit). [provided by Alliance of Genome Resources, Apr 2022] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
Rela
M08038_2.00 |
Mus musculus |
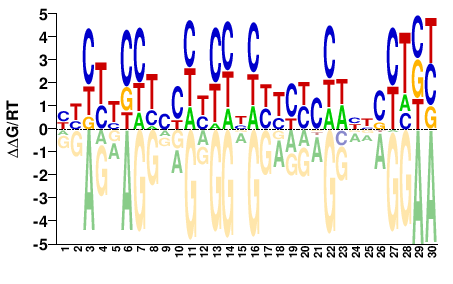
HHYYYCYYNYHYYYNHYYYYCYWNNSYTBY |
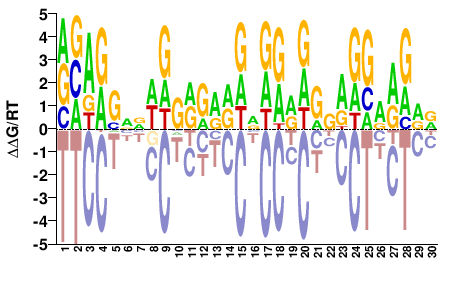
RVARSNNWRGRRRRDNRRRDRNRRGRRRDD |
ChIP-seq
Chen et al.(2011)
SRP001843_p65_Input |
(Direct) |
(Direct) |
Rela
M08039_2.00 |
Mus musculus |
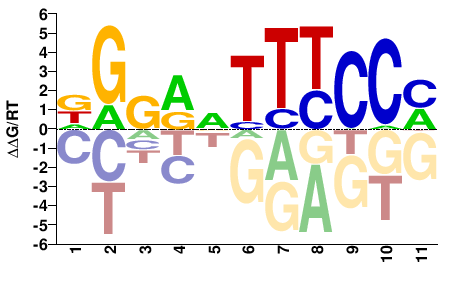
DGGRVTTTCCM |

KGGAAABYCCH |
ChIP-seq
Chen et al.(2011)
SRP001843_p65_Input_LPSstim |
(Direct) |
(Direct) |
Rela
M09355_2.00 |
Mus musculus |
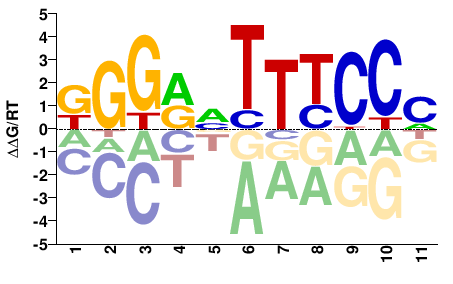
KGGRVTTYCCM |

KGGRAABYCCM |
Misc
Kulakovskiy et al.(2013)
TF65_MOUSE.H11MO.0.A |
(Direct) |
(Direct) |
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
RELA
M02711_2.00 |
Homo sapiens |
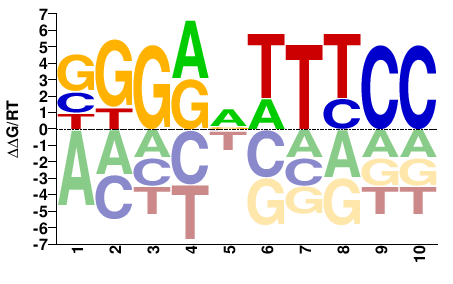
SGGRVTTTCC |

GGAAABYCCS |
SELEX
Mathelier et al.(2014)
MA0107.1
|
0.910
|
0.910
|
RELA
M07999_2.00 |
Homo sapiens |

DGGGRMTTTCCMVN |

NBKGGAAAKYCCCH |
ChIP-seq
Gerstein et al.(2012)
GM10847_NFKB_Stanford
|
0.910
|
0.910
|
RELA
M08000_2.00 |
Homo sapiens |
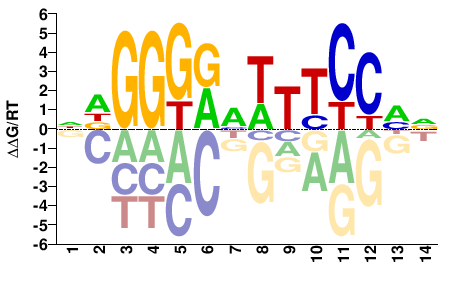
NDGGGRMWTTCCHN |
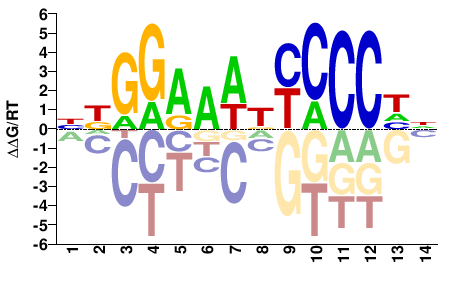
NDGGAAWKYCCCHN |
ChIP-seq
Gerstein et al.(2012)
GM12878_NFKB_Stanford
|
0.910
|
0.910
|
RELA
M08001_2.00 |
Homo sapiens |

NDGGGRMTTTCCMVN |

NBKGGAAAKYCCCHN |
ChIP-seq
Gerstein et al.(2012)
GM12891_NFKB_Stanford
|
0.910
|
0.910
|
RELA
M08002_2.00 |
Homo sapiens |

DGGGRHTTTCCMVNN |

NNBKGGAAADYCCCH |
ChIP-seq
Gerstein et al.(2012)
GM12892_NFKB_Stanford
|
0.910
|
0.910
|
RELA
M08003_2.00 |
Homo sapiens |

DGGGRVTTTCCHVN |

NBDGGAAABYCCCH |
ChIP-seq
Gerstein et al.(2012)
GM15510_NFKB_Stanford
|
0.910
|
0.910
|
RELA
M08004_2.00 |
Homo sapiens |

NDGGGRMTTTCCMNN |

NNKGGAAAKYCCCHN |
ChIP-seq
Gerstein et al.(2012)
GM18505_NFKB_Stanford
|
0.910
|
0.910
|
RELA
M08005_2.00 |
Homo sapiens |

NDGGGRHTTTCCMVN |
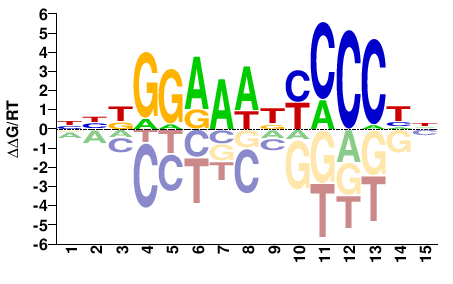
NBKGGAAADYCCCHN |
ChIP-seq
Gerstein et al.(2012)
GM18526_NFKB_Stanford
|
0.910
|
0.910
|
RELA
M08006_2.00 |
Homo sapiens |
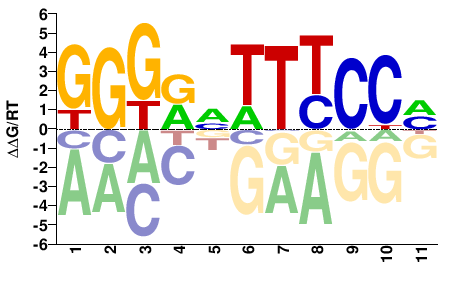

GGGRVTTTCCM |

KGGAAABYCCC |
ChIP-seq
Gerstein et al.(2012)
GM18951_NFKB_Stanford
|
0.910
|
0.910
|
RELA
M08007_2.00 |
Homo sapiens |

DGGGRMTTTCCMV |

BKGGAAAKYCCCH |
ChIP-seq
Gerstein et al.(2012)
GM19099_NFKB_Stanford
|
0.910
|
0.910
|
RELA
M08008_2.00 |
Homo sapiens |
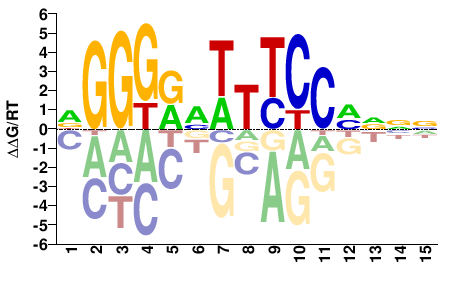

DGGGRMTTTCCMNNN |

NNNKGGAAAKYCCCH |
ChIP-seq
Gerstein et al.(2012)
GM19193_NFKB_Stanford
|
0.910
|
0.910
|
RELA
M09351_2.00 |
Homo sapiens |

NWGGGRHWTTCCMV |

BKGGAAWDYCCCWN |
Misc
Kulakovskiy et al.(2013)
TF65_HUMAN.H11MO.0.A
|
0.910
|
0.910
|
RELA
M09626_2.00 |
Homo sapiens |
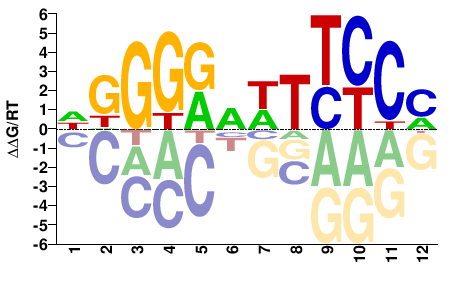
DGGGRVWTTCCM |

KGGAAWBYCCCH |
Misc
Heinz et al.(2010)
GM12787-p65_GSE19485
|
0.910
|
0.910
|
RELA
M11237_2.00 |
Homo sapiens |

BGGRVTTTCC |

GGAAABYCCV |
Transfac
Matys et al.(2006)
V$NFKAPPAB65_01
|
0.910
|
0.910
|
RELA
M11238_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$RELA_02
|
0.910
|
0.910
|
RELA
M11239_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$RELA_03
|
0.910
|
0.910
|
RELA
M11240_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$RELA_04
|
0.910
|
0.910
|
RELA
M11241_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$RELA_05
|
0.910
|
0.910
|
RELA
M11242_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$RELA_Q6
|
0.910
|
0.910
|
| For this family, TFs with SR scores > 0.700 will likely have a similar motif |