|
|
IKZF1
(Homo sapiens)
C2H2 ZF
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
Animal TF db |
| PF00096 (zf-C2H2) |
IPR007087 |
ENSG00000185811 |
T095237_2.00 |
Ensembl (2018-Dec-8) |
Link out |
| NCBI Gene Info:This gene encodes a transcription factor that belongs to the family of zinc-finger DNA-binding proteins associated with chromatin remodeling. The expression of this protein is restricted to the fetal and adult hemo-lymphopoietic system, and it functions as a regulator of lymphocyte differentiation. Several alternatively spliced transcript variants encoding different isoforms have been described for this gene. Most isoforms share a common C-terminal domain, which contains two zinc finger motifs that are required for hetero- or homo-dimerization, and for interactions with other proteins. The isoforms, however, differ in the number of N-terminal zinc finger motifs that bind DNA and in nuclear localization signal presence, resulting in members with and without DNA-binding properties. Only a few isoforms contain the requisite three or more N-terminal zinc motifs that confer high affinity binding to a specific core DNA sequence element in the promoters of target genes. The non-DNA-binding isoforms are largely found in the cytoplasm, and are thought to function as dominant-negative factors. Overexpression of some dominant-negative isoforms have been associated with B-cell malignancies, such as acute lymphoblastic leukemia (ALL). [provided by RefSeq, May 2014] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
IKZF1
M08939_2.00 |
Homo sapiens |
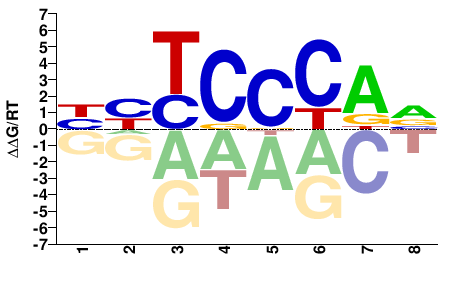
HYTCCCAV |
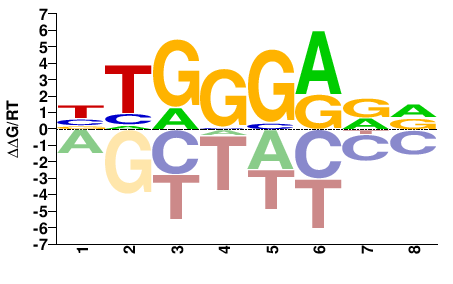
BTGGGARD |
Misc
Kulakovskiy et al.(2013)
IKZF1_HUMAN.H11MO.0.C |
(Direct) |
(Direct) |
IKZF1
M10301_2.00 |
Homo sapiens |

NHBYGGGAAYRYB |
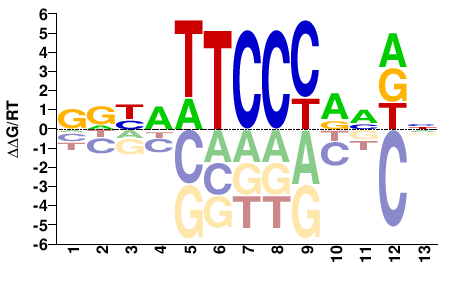
VRYRTTCCCRVDN |
Transfac
Matys et al.(2006)
V$IK1_01 |
(Direct) |
(Direct) |
IKZF1
M10302_2.00 |
Homo sapiens |
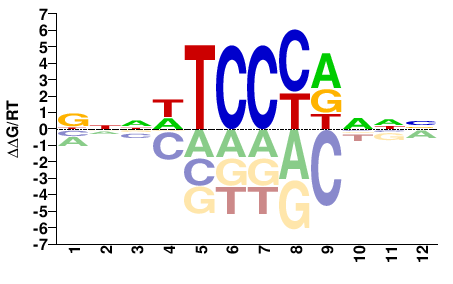
BNNWTCCCRNNN |
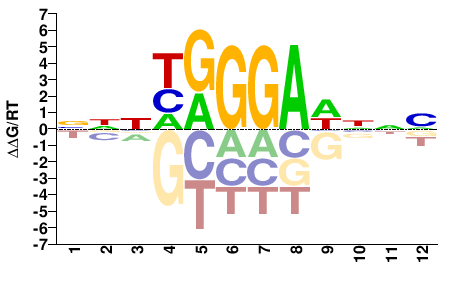
NNNYGGGAWNNV |
Transfac
Matys et al.(2006)
V$IK2_01 |
(Direct) |
(Direct) |
IKZF1
M10303_2.00 |
Homo sapiens |

KNYHGGGAAYRYY |

RRYRTTCCCDRNM |
Transfac
Matys et al.(2006)
V$IK3_01 |
(Direct) |
(Direct) |
IKZF1
M10304_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$IK_Q5_01 |
(Direct) |
(Direct) |
IKZF1
M10305_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$IK_Q5 |
(Direct) |
(Direct) |
IKZF1
M10306_2.00 |
Homo sapiens |

YYTCCCARA |
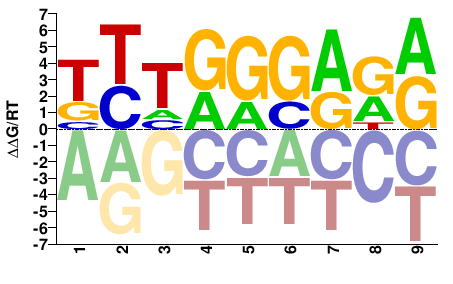
TYTGGGARR |
Transfac
Matys et al.(2006)
V$LYF1_01 |
(Direct) |
(Direct) |
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
Ikzf1
M08969_2.00 |
Mus musculus |
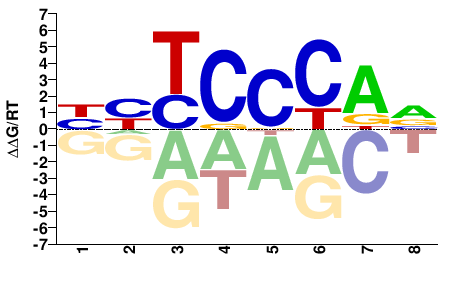
HYTCCCAV |
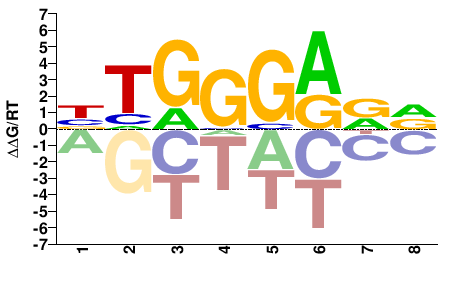
BTGGGARD |
Misc
Kulakovskiy et al.(2013)
IKZF1_MOUSE.H11MO.0.C
|
0.827
|
0.989
|
IKZF2
M10105_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$HELIOSA_01
|
0.804
|
0.913
|
IKZF2
M10106_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$HELIOSA_02
|
0.804
|
0.913
|
IKZF3
M08320_2.00 |
Homo sapiens |
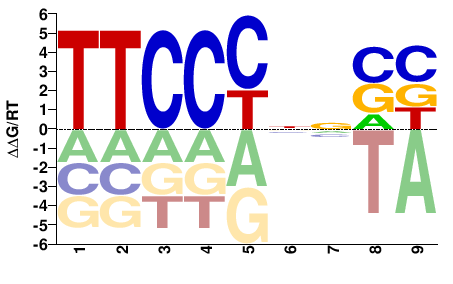
TTCCCNNSB |
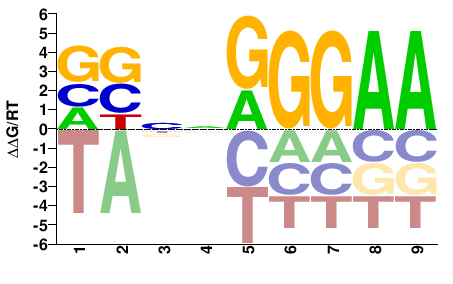
VSNNGGGAA |
ChIP-seq
Schmitges et al.(2016)
Q9UKT9_2-4.RCADE
|
0.790
|
0.870
|
| For this family, TFs with SR scores > 0.755 will likely have a similar motif |