|
|
ENSP00000265354
(Macaca fascicularis)
MADS box
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
| PF00319 (SRF-TF) |
IPR002100 |
ENSP00000265354 |
T256989_2.00 |
GigaDB (2015-Oct-22) |
| NCBI Gene Info:This gene encodes a ubiquitous nuclear protein that stimulates both cell proliferation and differentiation. It is a member of the MADS (MCM1, Agamous, Deficiens, and SRF) box superfamily of transcription factors. This protein binds to the serum response element (SRE) in the promoter region of target genes. This protein regulates the activity of many immediate-early genes, for example c-fos, and thereby participates in cell cycle regulation, apoptosis, cell growth, and cell differentiation. This gene is the downstream target of many pathways; for example, the mitogen-activated protein kinase pathway (MAPK) that acts through the ternary complex factors (TCFs). Two transcript variants encoding different isoforms have been found for this gene. [provided by RefSeq, May 2014] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
| No direct experiments |
|
|
|
|
|
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
srfA
M02283_2.00 |
Dictyostelium discoideum |

MYDWWWNNRD |
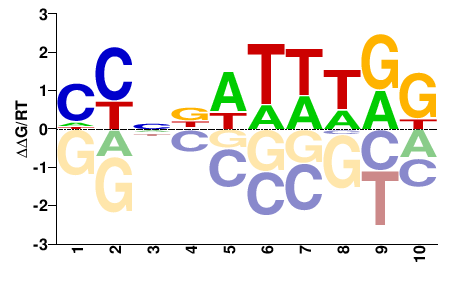
HYNNWWWHRK |
PBM
Weirauch et al.(2014)
pTH5539
|
0.867
|
0.867
|
Srf
M00178_2.00 |
Mus musculus |
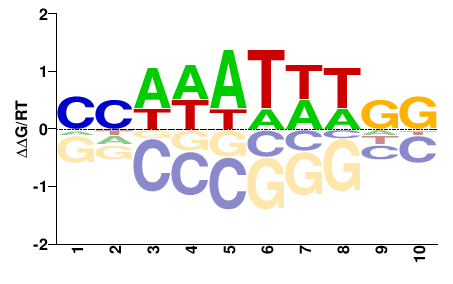
NNDDWWHHNN |
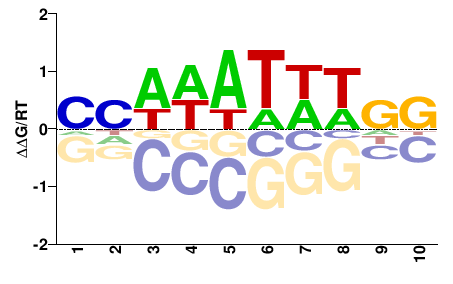
NNDDWWHHNN |
PBM
Badis et al.(2009)
Srf_3509
|
0.867
|
0.867
|
SRF
M03341_2.00 |
Homo sapiens |

DCCWTATATGGT |

ACCATATAWGGH |
SELEX
Jolma et al.(2013)
SRF_1
|
0.867
|
0.867
|
SRF
M03342_2.00 |
Homo sapiens |

TKHCCWTATATGGKMA |
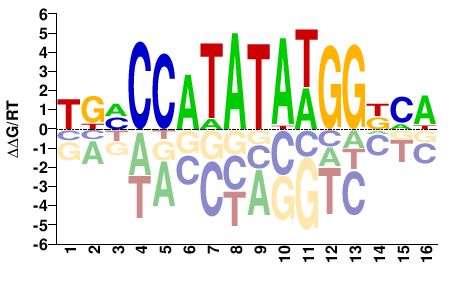
TKMCCATATAWGGDMA |
SELEX
Jolma et al.(2013)
SRF_2
|
0.867
|
0.867
|
SRF
M05553_2.00 |
Homo sapiens |
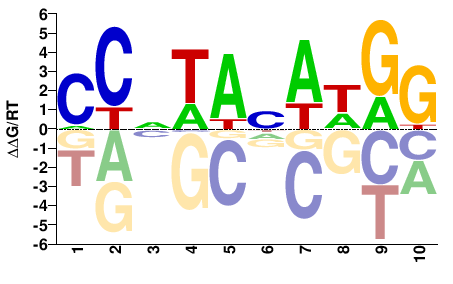
CCNTANAWGG |
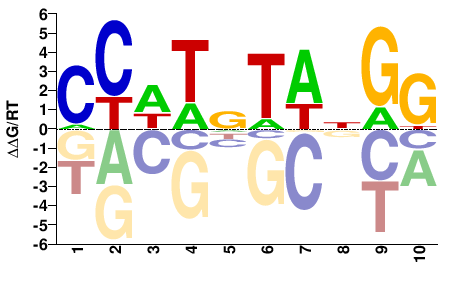
CCWTNTANGG |
SELEX
Yin et al.(2017)
SRF_eDBD_HT-SELEX
|
0.867
|
0.867
|
SRF
M07981_2.00 |
Homo sapiens |
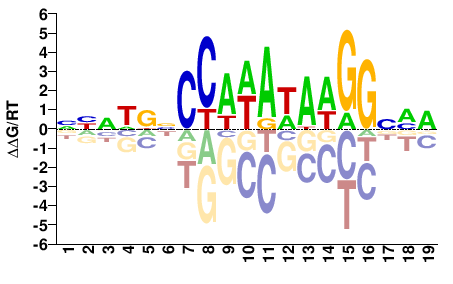
NNNWDNCCAWAWAWGGNVD |
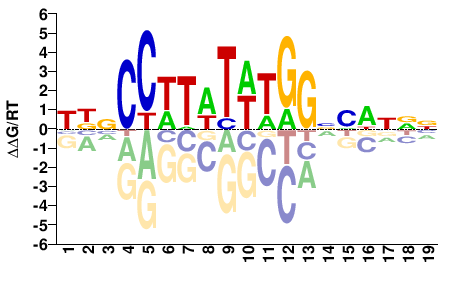
HBNCCWTWTWTGGNHWNNN |
ChIP-seq
Gerstein et al.(2012)
GM12878_SRF_HudsonAlpha
|
0.867
|
0.867
|
SRF
M07982_2.00 |
Homo sapiens |
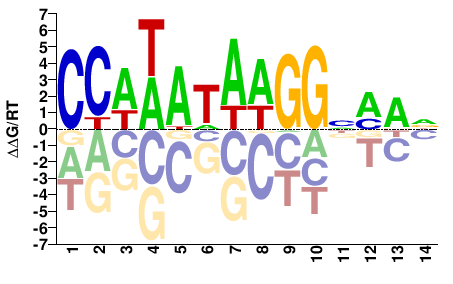
CCAWATAAGGNMAD |
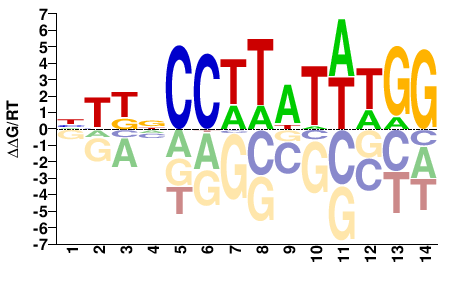
HTKNCCTTATWTGG |
ChIP-seq
Gerstein et al.(2012)
H1-hESC_SRF_HudsonAlpha
|
0.867
|
0.867
|
SRF
M07983_2.00 |
Homo sapiens |
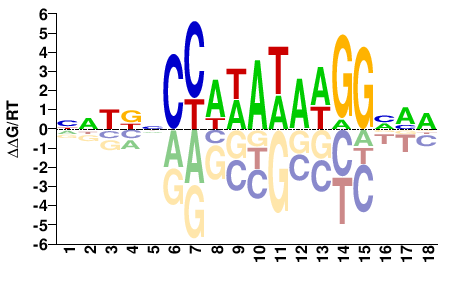
NNWNNCCAWAWAWGGNVD |
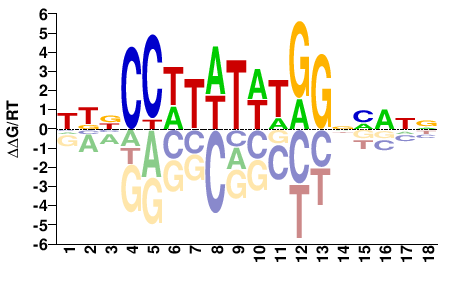
HBNCCWTWTWTGGNNWNN |
ChIP-seq
Gerstein et al.(2012)
HepG2_SRF_HudsonAlpha
|
0.867
|
0.867
|
SRF
M07984_2.00 |
Homo sapiens |
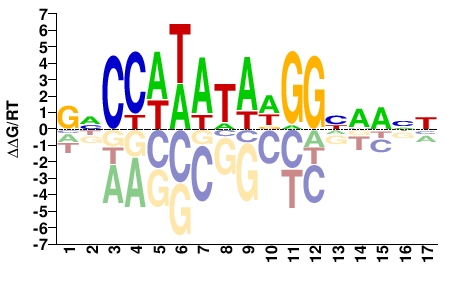
GNCCAWATADGGHMANN |
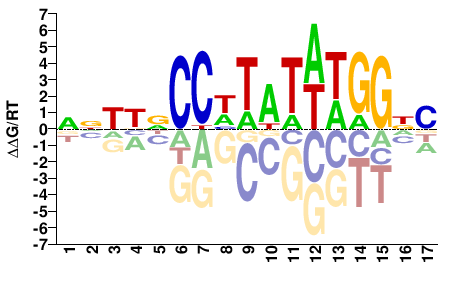
NNTKDCCHTATWTGGNC |
ChIP-seq
Gerstein et al.(2012)
K562_SRF_HudsonAlpha
|
0.867
|
0.867
|
SRF
M09249_2.00 |
Homo sapiens |
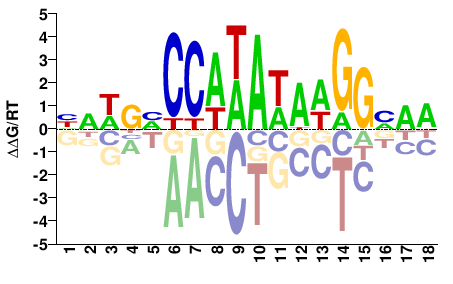
HNWBVCCAWAWAWGGNRR |
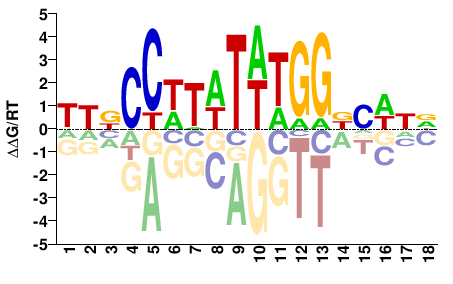
YYNCCWTWTWTGGBVWND |
Misc
Kulakovskiy et al.(2013)
SRF_HUMAN.H11MO.0.A
|
0.867
|
0.867
|
Srf
M09254_2.00 |
Mus musculus |
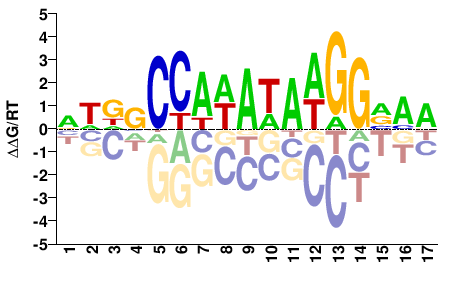
NWDVCCAWAWAWGGVMR |
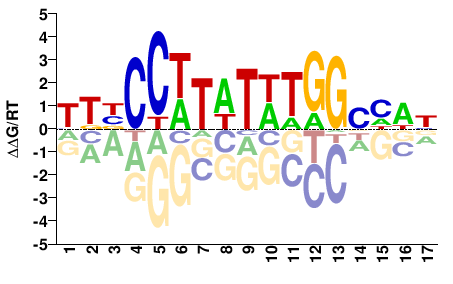

YKBCCWTWTWTGGBHWN |
Misc
Kulakovskiy et al.(2013)
SRF_MOUSE.H11MO.0.A
|
0.867
|
0.867
|
Srf
M09598_2.00 |
Mus musculus |

CCWWATWWGGNH |
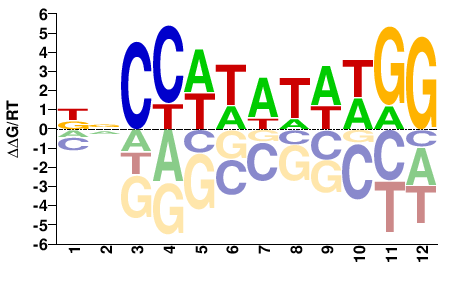
DNCCWWATWWGG |
Misc
Heinz et al.(2010)
PUER-Srf_Sullivan_et_al.
|
0.867
|
0.867
|
SRF
M10947_2.00 |
Homo sapiens |
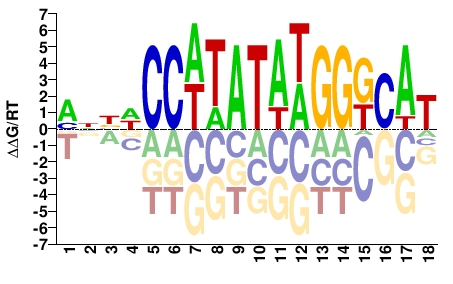
MNBWCCWTATAWGGGCAT |
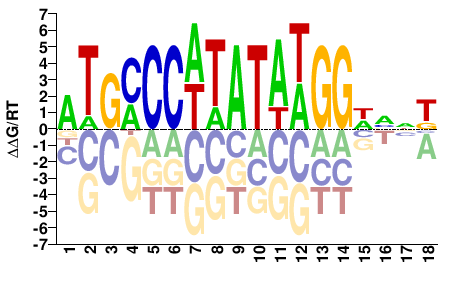
ATGCCCWTATAWGGWVNK |
Transfac
Matys et al.(2006)
V$SRF_01
|
0.867
|
0.867
|
SRF
M10948_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SRF_02
|
0.867
|
0.867
|
SRF
M10949_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SRF_03
|
0.867
|
0.867
|
SRF
M10950_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SRF_09
|
0.867
|
0.867
|
SRF
M10951_2.00 |
Homo sapiens |
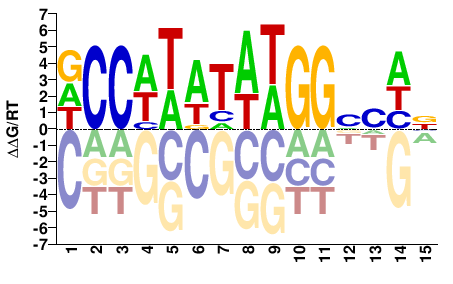
DCCWTATAWGGVSHB |
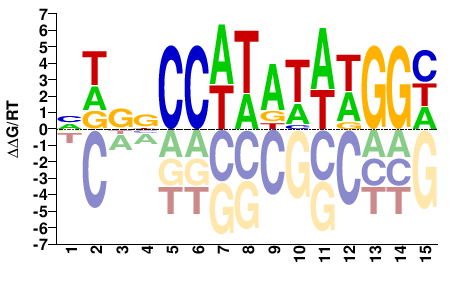
VDSBCCWTATAWGGH |
Transfac
Matys et al.(2006)
V$SRF_C
|
0.867
|
0.867
|
SRF
M10952_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SRF_Q3
|
0.867
|
0.867
|
SRF
M10953_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SRF_Q4
|
0.867
|
0.867
|
SRF
M10954_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SRF_Q5_01
|
0.867
|
0.867
|
SRF
M10955_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$SRF_Q5_02
|
0.867
|
0.867
|
SRF
M10956_2.00 |
Homo sapiens |

BNCCAWATAWGGVN |
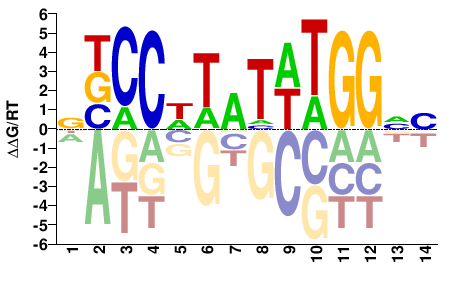
NBCCWTATWTGGNV |
Transfac
Matys et al.(2006)
V$SRF_Q6
|
0.867
|
0.867
|
SRF
M05554_2.00 |
Homo sapiens |

CCNTAYAWGG |
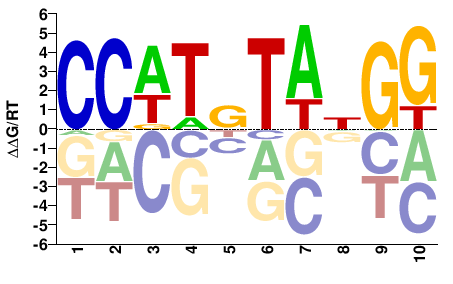

CCWTRTANGG |
SELEX
Yin et al.(2017)
SRF_eDBD_Methyl-HT-SELEX
|
0.867
|
0.867
|
bs
M03915_2.00 |
Drosophila melanogaster |

NCCWTATAWGGN |

NCCWTATAWGGN |
SELEX
Nitta et al.(2015)
bs_1
|
0.833
|
0.833
|
bs
M03916_2.00 |
Drosophila melanogaster |
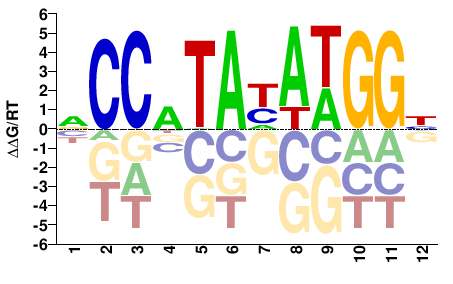
NCCWTAHAWGGH |

DCCWTDTAWGGN |
SELEX
Nitta et al.(2015)
bs_2
|
0.833
|
0.833
|
unc-120
M00684_2.00 |
Caenorhabditis elegans |

CHWAWWDGG |
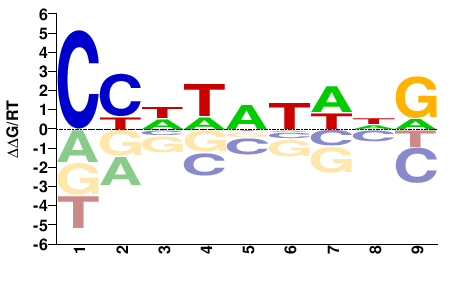
CCHWWTWDG |
PBM
Narasimhan et al.(2015)
pTH10822
|
0.767
|
0.767
|
MCM1
M01558_2.00 |
Saccharomyces cerevisiae |
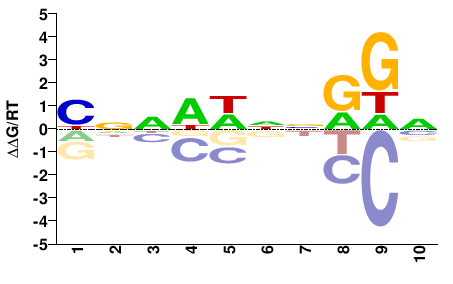
YNNWWNNRGN |
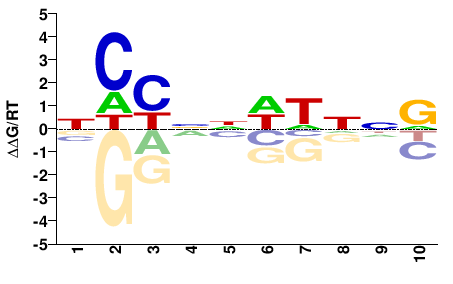
NCYNNWWNNR |
PBM
Zhu et al.(2009)
Mcm1
|
0.733
|
0.733
|
MCM1
M08589_2.00 |
Saccharomyces cerevisiae |
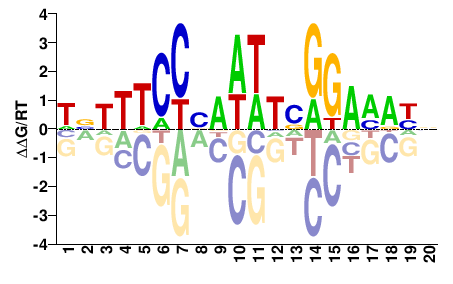

HNHTWCCBDWWHVGGAHDHN |
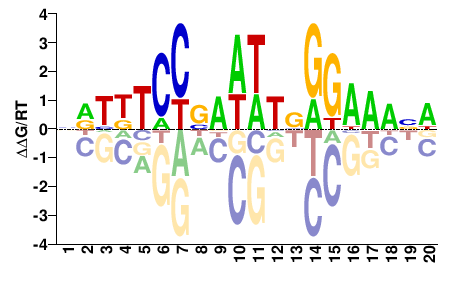

NDHDTCCBDWWHVGGWADND |
Misc
DeBoer et al.(2011)
YMR043W_831
|
0.733
|
0.733
|
MCM1
M10990_2.00 |
Saccharomyces cerevisiae |
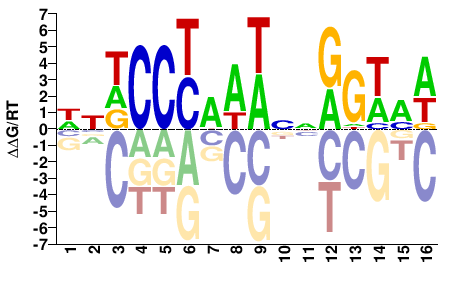
WNDCCYAAWNNGGWMW |
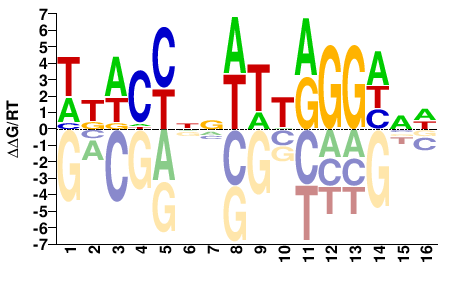
WKWCCNNWTTRGGHNW |
Transfac
Matys et al.(2006)
F$MCM1_01
|
0.733
|
0.733
|
MCM1
M10991_2.00 |
Saccharomyces cerevisiae |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
F$MCM1_02
|
0.733
|
0.733
|
MCM1
M10992_2.00 |
Saccharomyces cerevisiae |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
F$MCM1_Q6
|
0.733
|
0.733
|
MCM1
M08458_2.00 |
Saccharomyces cerevisiae |
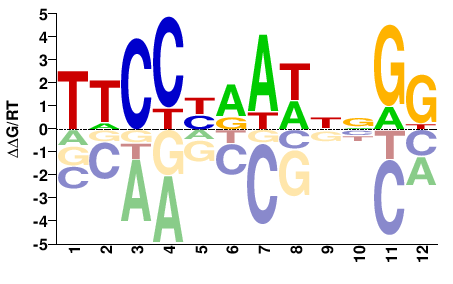

TTCCYRAWNNGG |

CCNNWTYRGGAA |
COMPILED
Mathelier et al.(2014)
MA0331.1
|
0.733
|
0.733
|
| For this family, TFs with SR scores > 0.700 will likely have a similar motif |