|
|
ENSP00000366022-D1
(Bos grunniens)
RFX
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
| PF02257 (RFX_DNA_binding) |
IPR003150 |
ENSP00000366022-D1 |
T318981_2.00 |
GigaDB (2015-Oct-22) |
| NCBI Gene Info:This gene encodes a tumor suppressor protein containing transcriptional activation, DNA binding, and oligomerization domains. The encoded protein responds to diverse cellular stresses to regulate expression of target genes, thereby inducing cell cycle arrest, apoptosis, senescence, DNA repair, or changes in metabolism. Mutations in this gene are associated with a variety of human cancers, including hereditary cancers such as Li-Fraumeni syndrome. Alternative splicing of this gene and the use of alternate promoters result in multiple transcript variants and isoforms. Additional isoforms have also been shown to result from the use of alternate translation initiation codons from identical transcript variants (PMIDs: 12032546, 20937277). [provided by RefSeq, Dec 2016] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
| No direct experiments |
|
|
|
|
|
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
PHUM046580
M02454_2.00 |
Pediculus humanus |

NGTHDCYANV |
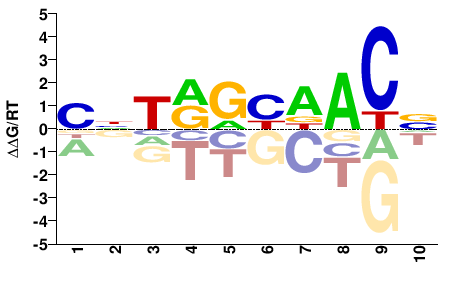
BNTRGHDACN |
PBM
Weirauch et al.(2014)
pTH9278
|
0.666
|
0.500
|
G4VLF2_SCHMA
M02452_2.00 |
Schistosoma mansoni |

GTTGCTAD |
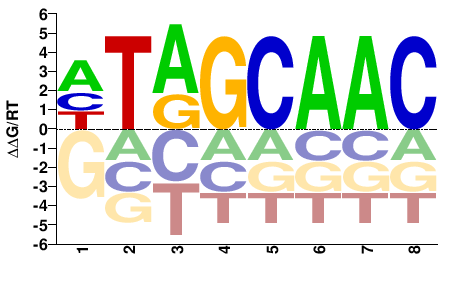
HTAGCAAC |
PBM
Weirauch et al.(2014)
pTH8587
|
0.662
|
0.514
|
RFX1
M02447_2.00 |
Anolis carolinensis |

GTHRCYANGN |
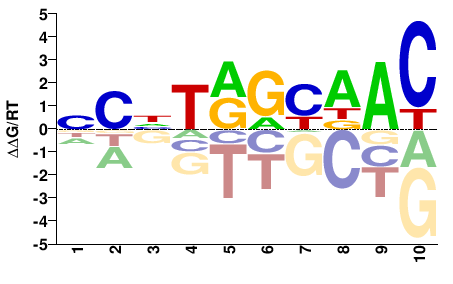
NCNTRGYDAC |
PBM
Weirauch et al.(2014)
pTH9223
|
0.662
|
0.486
|
RFX1
M02451_2.00 |
Tetraodon nigroviridis |
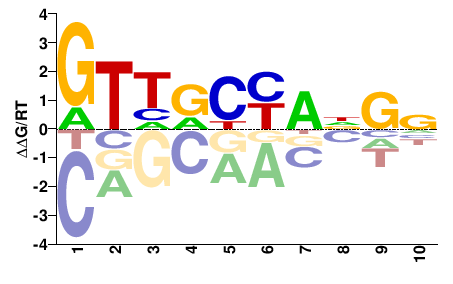
GTHDYYANGN |

NCNTRRHDAC |
PBM
Weirauch et al.(2014)
pTH9385
|
0.662
|
0.486
|
RFX1
M05723_2.00 |
Homo sapiens |
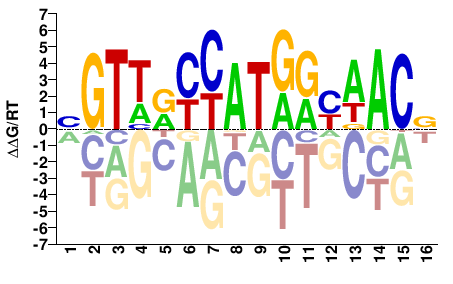
BGTTRYCATGRYAACV |
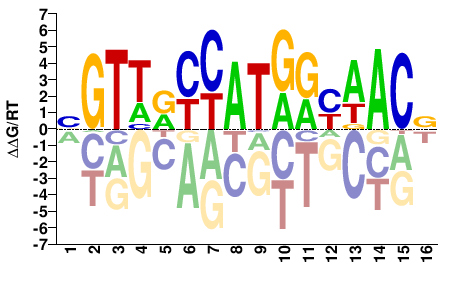
BGTTRYCATGRYAACV |
SELEX
Yin et al.(2017)
RFX1_eDBD_HT-SELEX
|
0.662
|
0.486
|
Rfx1
M08162_2.00 |
Mus musculus |
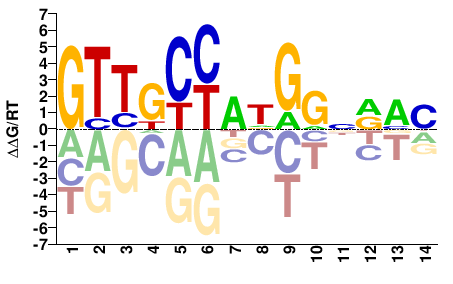
GTTGCYADGGNRAC |
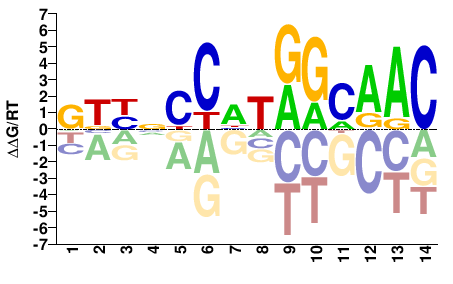

GTYNCCHTRGCAAC |
ChIP-seq
Mathelier et al.(2014)
MA0509.1
|
0.662
|
0.486
|
RFX1
M09363_2.00 |
Homo sapiens |
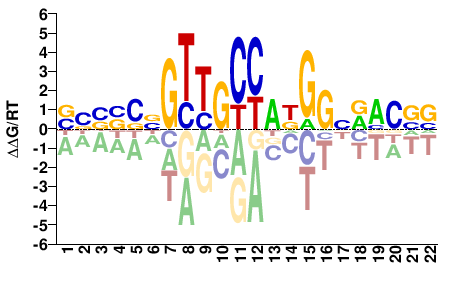
BBBSSNGTTGCCADGGNRMCVV |
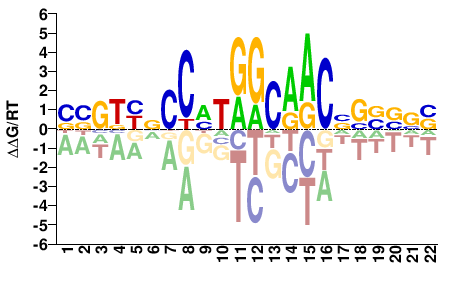

BBGKYNCCHTGGCAACNSSVVV |
Misc
Kulakovskiy et al.(2013)
RFX1_HUMAN.H11MO.0.B
|
0.662
|
0.486
|
Rfx1
M09367_2.00 |
Mus musculus |
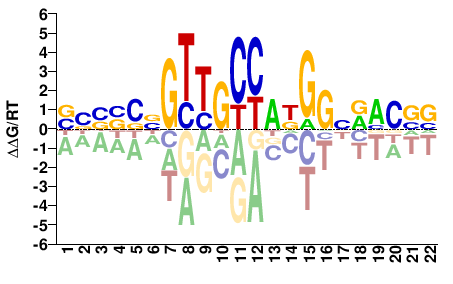
BBBSSNGTTGCCADGGNRMCVV |
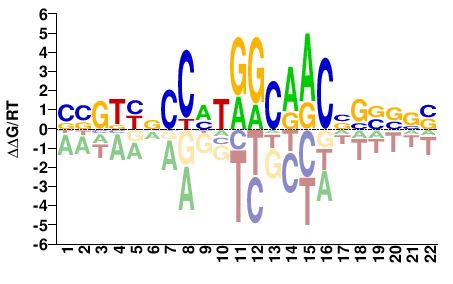

BBGKYNCCHTGGCAACNSSVVV |
Misc
Kulakovskiy et al.(2013)
RFX1_MOUSE.H11MO.0.A
|
0.662
|
0.486
|
RFX1
M11245_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$EFC_Q6
|
0.662
|
0.486
|
RFX1
M11246_2.00 |
Homo sapiens |

HNGTTRCYWVGYDMYNN |
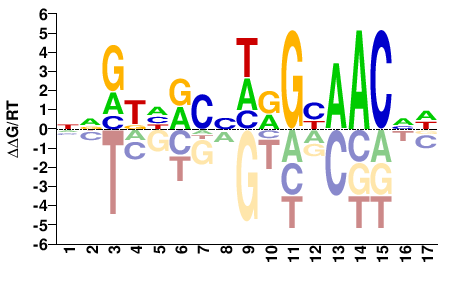
NNRKHRCBWRGYAACND |
Transfac
Matys et al.(2006)
V$RFX1_01
|
0.662
|
0.486
|
RFX1
M11247_2.00 |
Homo sapiens |

HNGTTRCYWBVGYDMYNN |
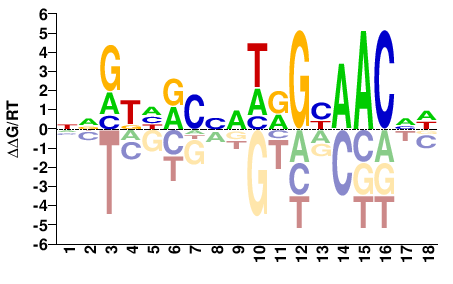
NNRKHRCBVWRGYAACND |
Transfac
Matys et al.(2006)
V$RFX1_02
|
0.662
|
0.486
|
RFX1
M11248_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$RFX1_Q6
|
0.662
|
0.486
|
RFX1
M05724_2.00 |
Homo sapiens |
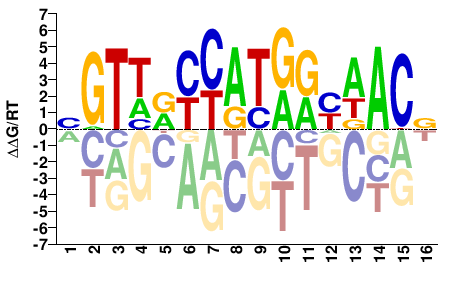
BGTWRYCATGRYWACV |
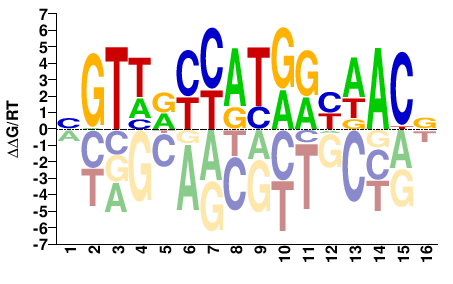
BGTWRYCATGRYWACV |
SELEX
Yin et al.(2017)
RFX1_eDBD_Methyl-HT-SELEX
|
0.662
|
0.486
|
Rfx3
M00188_2.00 |
Mus musculus |
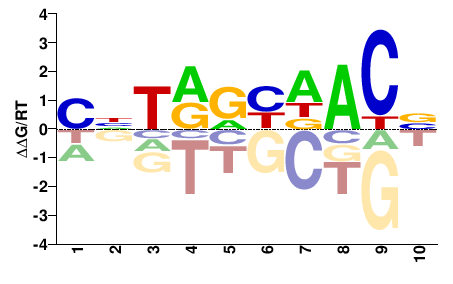
SNTRRHDACN |

NGTHDYYANS |
PBM
Badis et al.(2009)
Rfx3_3961
|
0.662
|
0.472
|
RFX3
M03452_2.00 |
Homo sapiens |
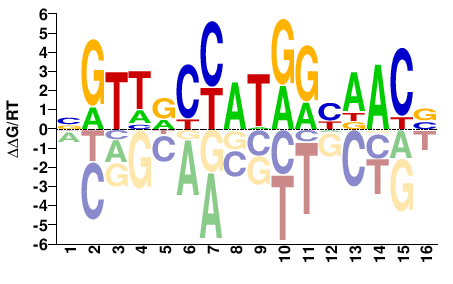
NGTTRCCATGGHAACV |
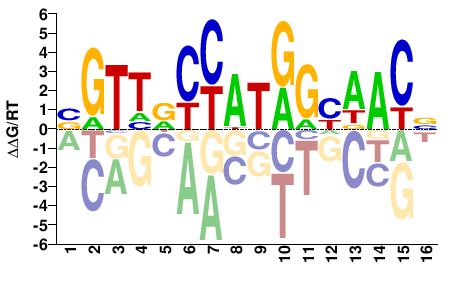
BGTTDCCATGGYAACN |
SELEX
Jolma et al.(2013)
RFX3_1
|
0.662
|
0.472
|
RFX3
M03453_2.00 |
Homo sapiens |
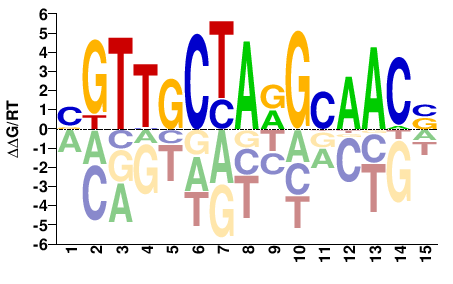
BGTTGCTARGCAACS |

SGTTGCYTAGCAACV |
SELEX
Jolma et al.(2013)
RFX3_2
|
0.662
|
0.472
|
Rfx3
M03463_2.00 |
Mus musculus |

SGTTRCCATGGYAACS |

SGTTRCCATGGYAACS |
SELEX
Jolma et al.(2013)
Rfx3_1
|
0.662
|
0.472
|
RFX3
M02750_2.00 |
Homo sapiens |

GTTRYYATGG |

CCATRRYAAC |
SELEX
Jolma et al.(2010)
RFX3_dimer
|
0.662
|
0.472
|
RFX3
M05715_2.00 |
Homo sapiens |
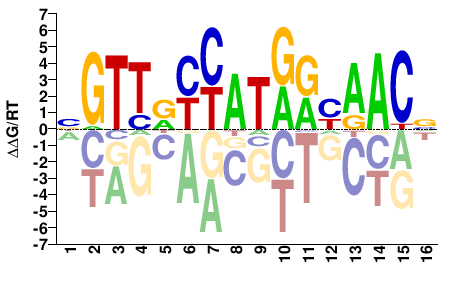
NGTTRYYATRRYAACN |
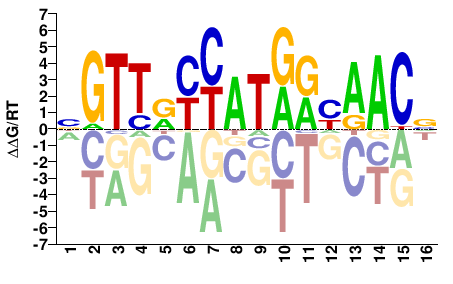
NGTTRYYATRRYAACN |
SELEX
Yin et al.(2017)
RFX3_eDBD_HT-SELEX_1
|
0.662
|
0.472
|
RFX3
M05716_2.00 |
Homo sapiens |

BGTTGCTARGCAACS |

SGTTGCYTAGCAACV |
SELEX
Yin et al.(2017)
RFX3_eDBD_HT-SELEX_2
|
0.662
|
0.472
|
RFX3
M09361_2.00 |
Homo sapiens |
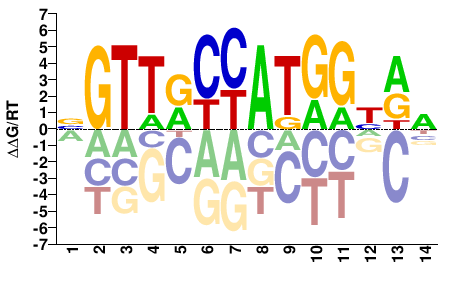
BGTTRCCATGGYRN |
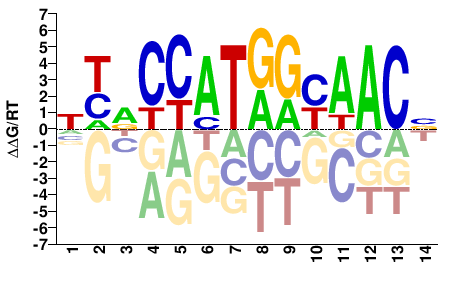
NYRCCATGGYAACV |
Misc
Kulakovskiy et al.(2013)
RFX3_HUMAN.H11MO.0.B
|
0.662
|
0.472
|
Rfx3
M09368_2.00 |
Mus musculus |
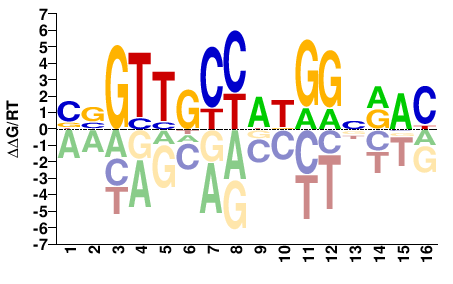
BBGTTGCCATGGNRAC |
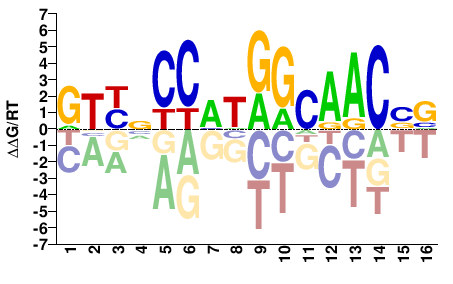

GTYNCCATGGCAACVV |
Misc
Kulakovskiy et al.(2013)
RFX3_MOUSE.H11MO.0.C
|
0.662
|
0.472
|
RFX3
M09627_2.00 |
Homo sapiens |
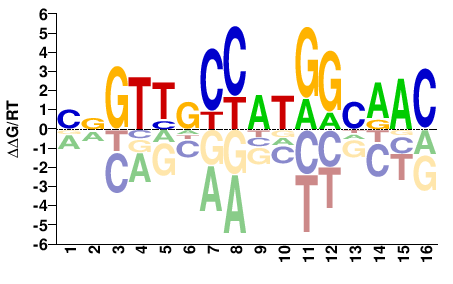
BNGTTGCCATGGCAAC |

GTTGCCATGGCAACNV |
Misc
Heinz et al.(2010)
K562-RFX3_SRA012198
|
0.662
|
0.472
|
RFX3
M05717_2.00 |
Homo sapiens |
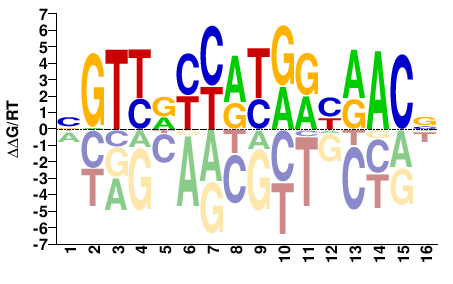
BGTTRYYATRRYAACV |
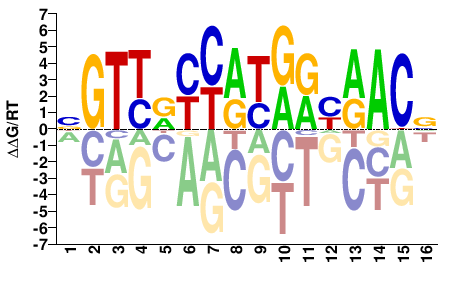
BGTTRYYATRRYAACV |
SELEX
Yin et al.(2017)
RFX3_eDBD_Methyl-HT-SELEX_1
|
0.662
|
0.472
|
RFX3
M05718_2.00 |
Homo sapiens |
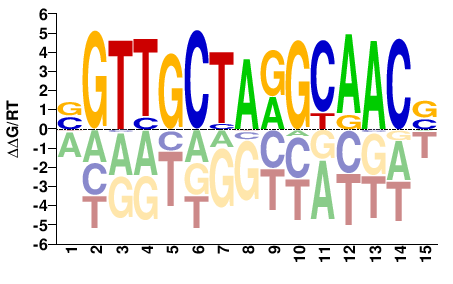
BGTTGCTARGCAACV |

BGTTGCYTAGCAACV |
SELEX
Yin et al.(2017)
RFX3_eDBD_Methyl-HT-SELEX_2
|
0.662
|
0.472
|
RFX2
M02449_2.00 |
Homo sapiens |

BGTHRYYADG |
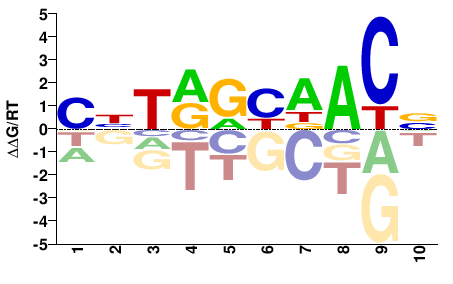
CHTRRYDACV |
PBM
Weirauch et al.(2014)
pTH9194
|
0.661
|
0.486
|
RFX2
M02450_2.00 |
Ornithorhynchus anatinus |

NGTHRYYANV |
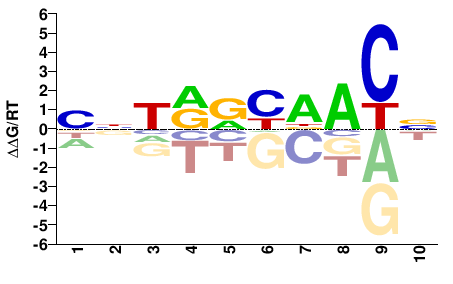
BNTRRYDACN |
PBM
Weirauch et al.(2014)
pTH9269
|
0.661
|
0.486
|
RFX2
M03454_2.00 |
Homo sapiens |
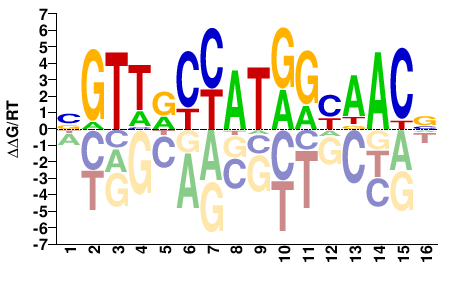
BGTTRCCATGGYAACV |
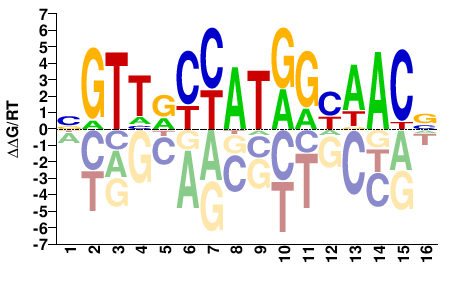
BGTTRCCATGGYAACV |
SELEX
Jolma et al.(2013)
RFX2_1
|
0.661
|
0.486
|
RFX2
M03455_2.00 |
Homo sapiens |
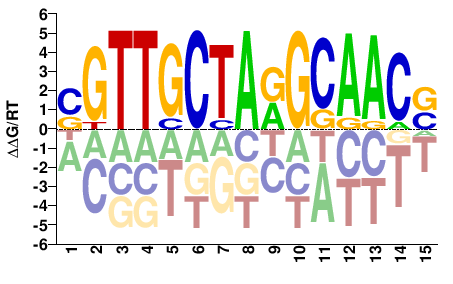
SGTTGCTARGCAACS |
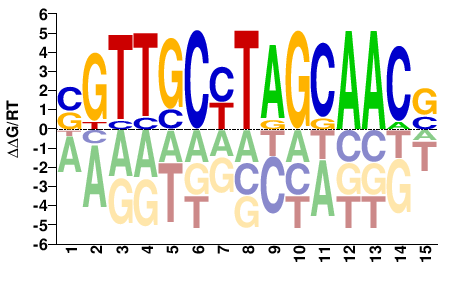
SGTTGCYTAGCAACS |
SELEX
Jolma et al.(2013)
RFX2_2
|
0.661
|
0.486
|
RFX2
M05719_2.00 |
Homo sapiens |

NGTWDYYATRRHWACN |

NGTWDYYATRRHWACN |
SELEX
Yin et al.(2017)
RFX2_eDBD_HT-SELEX
|
0.661
|
0.486
|
RFX2
M09362_2.00 |
Homo sapiens |
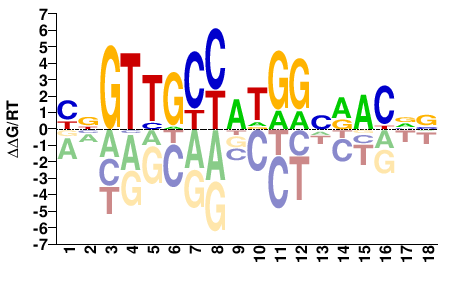
YNGTTGCCATGGVRACVV |
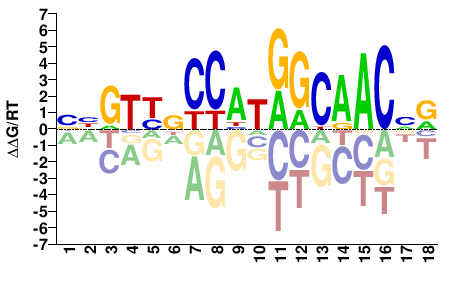
BBGTYBCCATGGCAACNR |
Misc
Kulakovskiy et al.(2013)
RFX2_HUMAN.H11MO.0.A
|
0.661
|
0.486
|
RFX2
M09628_2.00 |
Homo sapiens |

BGTTKCCATGGMAAC |

GTTKCCATGGMAACV |
Misc
Heinz et al.(2010)
LoVo-RFX2_GSE49402
|
0.661
|
0.486
|
RFX2
M05720_2.00 |
Homo sapiens |
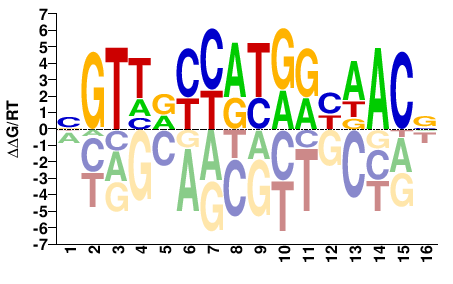
BGTTRCCATGRYAACV |
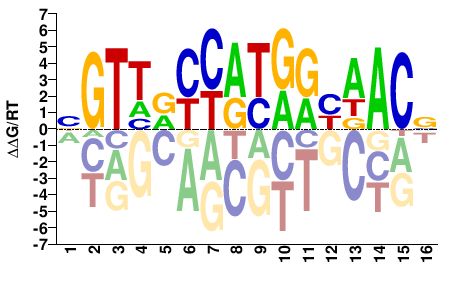
BGTTRYCATGGYAACV |
SELEX
Yin et al.(2017)
RFX2_eDBD_Methyl-HT-SELEX
|
0.661
|
0.486
|
daf-19
M00709_2.00 |
Caenorhabditis elegans |

SGTTRCTADG |

CHTAGYAACS |
PBM
Narasimhan et al.(2015)
pTH10021
|
0.661
|
0.486
|
Rfx6
M09365_2.00 |
Mus musculus |

BDGTWKCYNDG |

CHNRGMAACHV |
Misc
Kulakovskiy et al.(2013)
RFX6_MOUSE.H11MO.0.C
|
0.661
|
0.472
|
Sjc_0102030
M02455_2.00 |
Schistosoma japonicum |

GTTRCYANG |
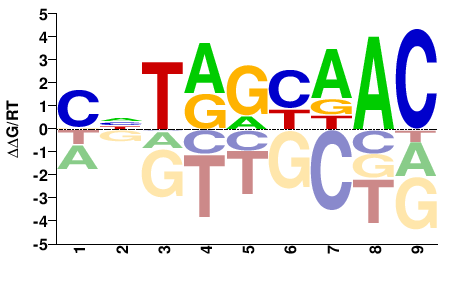
CNTRGYAAC |
PBM
Weirauch et al.(2014)
pTH9285
|
0.661
|
0.500
|
Rfx2
M03461_2.00 |
Mus musculus |

BGTTRCCATGGYAACV |

BGTTRCCATGGYAACV |
SELEX
Jolma et al.(2013)
Rfx2_1
|
0.661
|
0.486
|
Rfx2
M03462_2.00 |
Mus musculus |

SGTTGCTARGCAACV |

BGTTGCYTAGCAACS |
SELEX
Jolma et al.(2013)
Rfx2_2
|
0.661
|
0.486
|
Rfx2
M05888_2.00 |
Mus musculus |

NGTTRCYATGGHDAY |
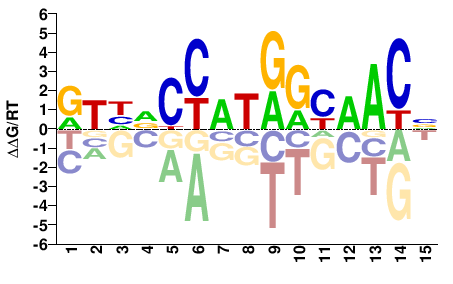
RTHDCCATRGYAACN |
SMiLE-seq
Isakova et al.(2017)
RFX2
|
0.661
|
0.486
|
Rfx2
M09366_2.00 |
Mus musculus |
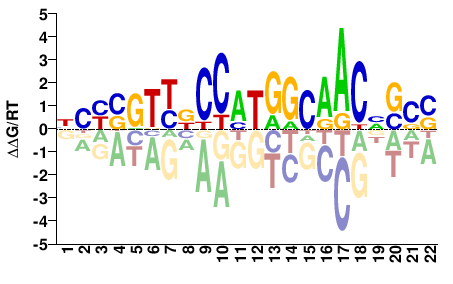
NBYBGTYNCCMTGGCAACNSSS |
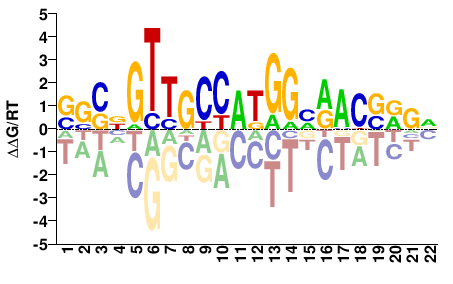
SSSNGTTGCCAKGGNRACVRVN |
Misc
Kulakovskiy et al.(2013)
RFX2_MOUSE.H11MO.0.A
|
0.661
|
0.486
|
Rfx4
M00186_2.00 |
Mus musculus |
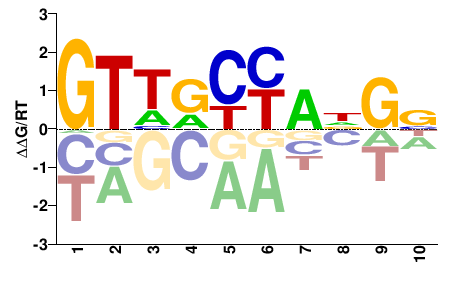
GTHDYYNNSN |

NSNNRRHDAC |
PBM
Badis et al.(2009)
Rfx4_3761
|
0.660
|
0.472
|
RFX4
M03456_2.00 |
Homo sapiens |
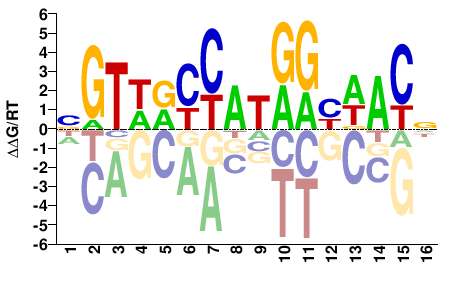
NGTWRYCATRGHWACN |
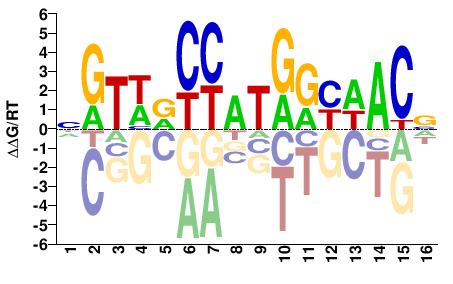
NGTWDCYATGRYWACN |
SELEX
Jolma et al.(2013)
RFX4_1
|
0.660
|
0.472
|
RFX4
M03457_2.00 |
Homo sapiens |

NGTTGCYRRGCAACS |

SGTTGCYYRGCAACN |
SELEX
Jolma et al.(2013)
RFX4_2
|
0.660
|
0.472
|
RFX4
M05721_2.00 |
Homo sapiens |
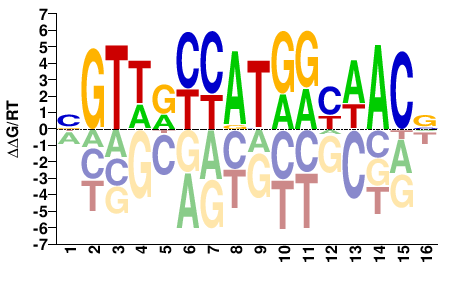
BGTWRYCATGRYWACV |
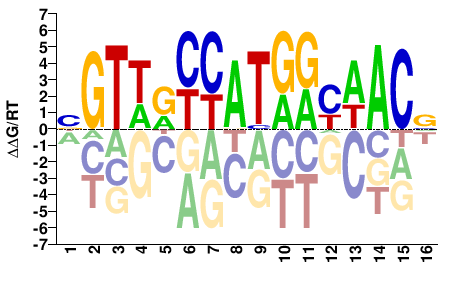
BGTWRYCATGRYWACV |
SELEX
Yin et al.(2017)
RFX4_eDBD_HT-SELEX
|
0.660
|
0.472
|
RFX4
M05722_2.00 |
Homo sapiens |
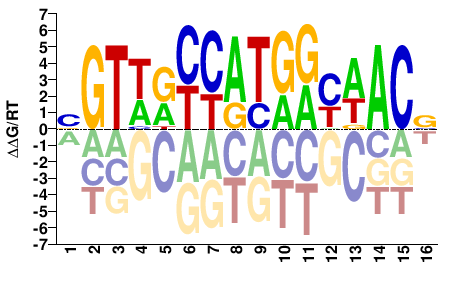
BGTWRYCATGRYWACV |
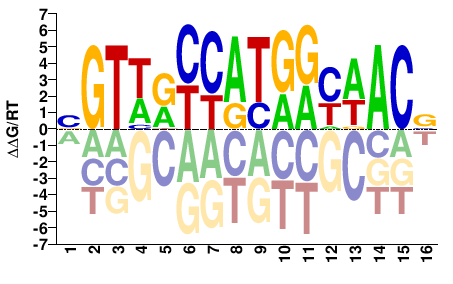
BGTWRYCATGRYWACV |
SELEX
Yin et al.(2017)
RFX4_eDBD_Methyl-HT-SELEX
|
0.660
|
0.472
|
RFX4
M02448_2.00 |
Cavia porcellus |
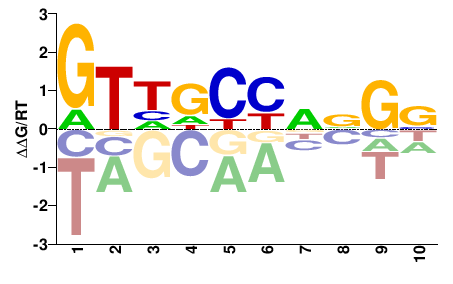
GTHDYYNNGN |
NCNNRRHDAC |
PBM
Weirauch et al.(2014)
pTH9249
|
0.660
|
0.458
|
XP_002157607.1
M02456_2.00 |
Hydra magnipapillata |
GTHDYYDNVN |

NBNHRRHDAC |
PBM
Weirauch et al.(2014)
pTH9199
|
0.659
|
0.458
|
GB41876
M02453_2.00 |
Apis mellifera |
GTHDCYANVN |
NBNTRGHDAC |
PBM
Weirauch et al.(2014)
pTH9226
|
0.658
|
0.472
|
Rfx
M03959_2.00 |
Drosophila melanogaster |
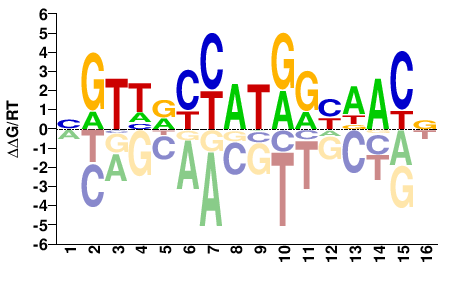
NGTHRCYATRGYDACN |
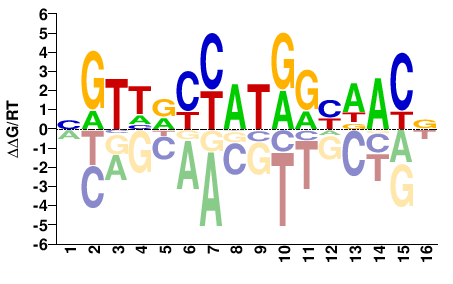
NGTHRCYATRGYDACN |
SELEX
Nitta et al.(2015)
Rfx_1
|
0.655
|
0.472
|
Rfx
M03960_2.00 |
Drosophila melanogaster |

GTTGCTARGCAAC |

GTTGCYTAGCAAC |
SELEX
Nitta et al.(2015)
Rfx_2
|
0.655
|
0.472
|
Rfx
M03961_2.00 |
Drosophila melanogaster |
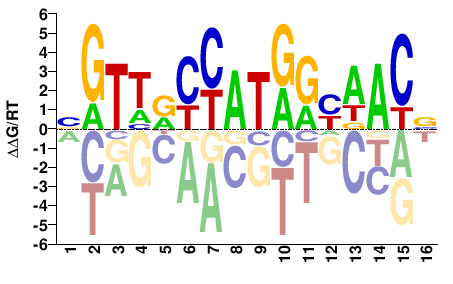
BGTWRCCATGGYWACV |
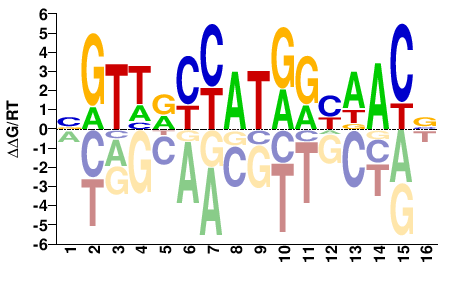
BGTWRCCATGGYWACV |
SELEX
Nitta et al.(2015)
Rfx_3
|
0.655
|
0.472
|
Rfx
M03962_2.00 |
Drosophila melanogaster |
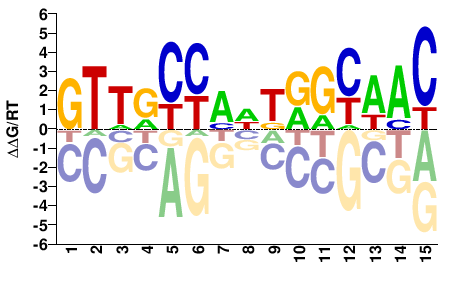
GTTRCYAWTRGYAAC |
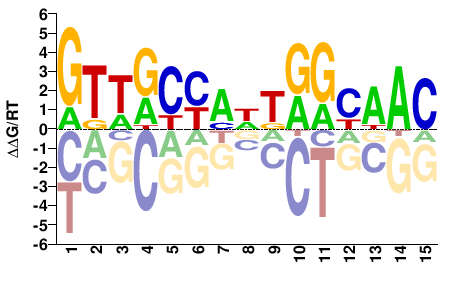
GTTRCYAWTRGYAAC |
SELEX
Nitta et al.(2015)
Rfx_4
|
0.655
|
0.472
|
Rfx
M03963_2.00 |
Drosophila melanogaster |

GTTGCTARGCAAC |
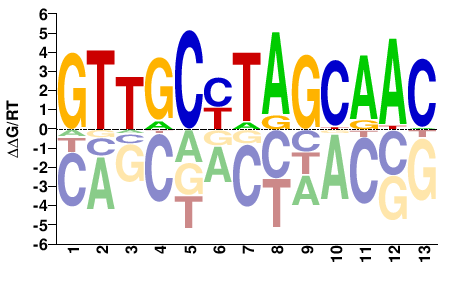

GTTGCYTAGCAAC |
SELEX
Nitta et al.(2015)
Rfx_5
|
0.655
|
0.472
|
| For this family, TFs with SR scores > 0.650 will likely have a similar motif |