|
|
Myc
(Rattus norvegicus)
bHLH
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
Animal TF db |
| PF00010 (HLH) |
IPR001092 |
ENSRNOG00000004500 |
T037175_2.00 |
Ensembl (2018-Dec-8) |
Link out |
| NCBI Gene Info:The protein encoded by this gene is a multifunctional, nuclear phosphoprotein that plays a role in cell cycle progression, apoptosis and cellular transformation. It functions as a transcription factor that regulates transcription of specific target genes. Mutations, overexpression, rearrangement and translocation of this gene have been associated with a variety of hematopoietic tumors, leukemias and lymphomas, including Burkitt lymphoma, in human. There is evidence to show that alternative translation initiations from an upstream, in-frame non-AUG (CUG) and a downstream AUG start site result in the production of two isoforms with distinct N-termini, in human and mouse. Rat mRNA also has a similarly placed CUG upstream of the AUG start site, suggesting that it may also produce two Myc proteins. [provided by RefSeq, Jul 2008] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
| No direct experiments |
|
|
|
|
|
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
Myc
M08751_2.00 |
Mus musculus |

GVSCACGTGGNN |

NNCCACGTGSBC |
Misc
Kulakovskiy et al.(2013)
MYC_MOUSE.H11MO.0.A
|
0.898
|
0.987
|
Myc
M09467_2.00 |
Mus musculus |

SCACGTGSNN |

NNSCACGTGS |
Misc
Heinz et al.(2010)
mES-cMyc_GSE11431
|
0.898
|
0.987
|
MYC
M08054_2.00 |
Homo sapiens |

NNSCACGTGGNN |
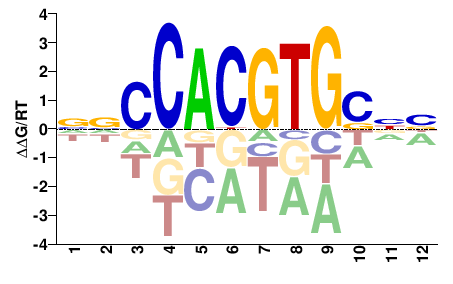

NNCCACGTGSNN |
ChIP-seq
Mathelier et al.(2014)
MA0147.3
|
0.846
|
0.960
|
MYC
M07799_2.00 |
Homo sapiens |
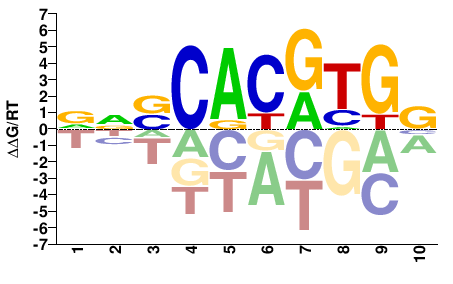
VNSCACGTGG |

CCACGTGSNB |
ChIP-seq
Gerstein et al.(2012)
HeLa-S3_CMYC_Stanford
|
0.846
|
0.960
|
MYC
M07800_2.00 |
Homo sapiens |

NNSCACGTGGB |
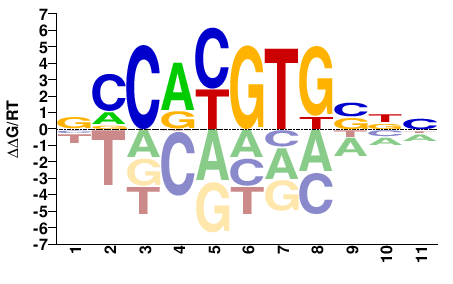

VCCACGTGSNN |
ChIP-seq
Gerstein et al.(2012)
HepG2_CMYC_UT-A
|
0.846
|
0.960
|
MYC
M07801_2.00 |
Homo sapiens |

VNVCACGTGGN |
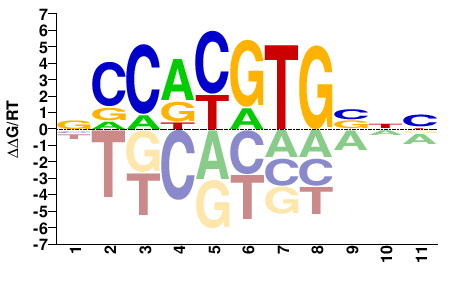
NCCACGTGBNB |
ChIP-seq
Gerstein et al.(2012)
K562_CMYC_Stanford
|
0.846
|
0.960
|
MYC
M07802_2.00 |
Homo sapiens |

VSCRCGTGBNB |

VNVCACGYGSB |
ChIP-seq
Gerstein et al.(2012)
K562_CMYC_UT-A
|
0.846
|
0.960
|
MYC
M07803_2.00 |
Homo sapiens |

CACGTGSH |

DSCACGTG |
ChIP-seq
Gerstein et al.(2012)
MCF-7_CMYC_UT-A
|
0.846
|
0.960
|
MYC
M07804_2.00 |
Homo sapiens |

NSCACGTGKBN |
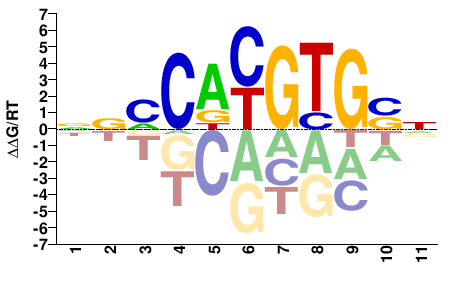

NVMCACGTGSN |
ChIP-seq
Gerstein et al.(2012)
NB4_CMYC_Stanford
|
0.846
|
0.960
|
MYC
M08729_2.00 |
Homo sapiens |
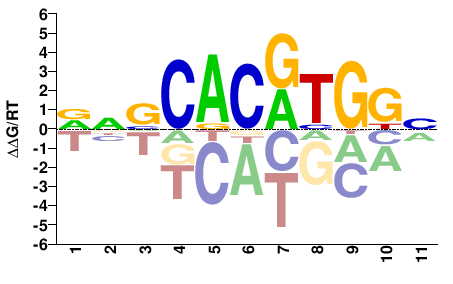
VNVCACRTGGN |
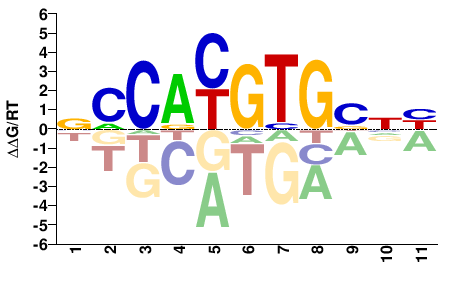

NCCAYGTGBNB |
Misc
Kulakovskiy et al.(2013)
MYC_HUMAN.H11MO.0.A
|
0.846
|
0.960
|
MYC
M09457_2.00 |
Homo sapiens |

CACGTGSN |
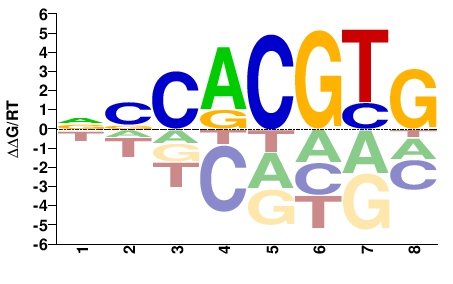
NSCACGTG |
Misc
Heinz et al.(2010)
LNCAP-cMyc_Unpublished
|
0.846
|
0.960
|
MYC
M09850_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$CMYC_01
|
0.846
|
0.960
|
MYC
M09851_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$CMYC_02
|
0.846
|
0.960
|
MYC
M09852_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$CMYC_Q3
|
0.846
|
0.960
|
MYC
M09853_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$CMYC_Q6_01
|
0.846
|
0.960
|
MYC
M09854_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$MYC_02
|
0.846
|
0.960
|
| For this family, TFs with SR scores > 0.838 will likely have a similar motif |