|
|
ENSP00000356959
(Macaca fascicularis)
Nuclear receptor
|
TF Information |
|---|
| Pfam ID |
Interpro ID |
Gene ID |
CIS-BP ID |
Sequence source |
| PF00105 (zf-C4) |
IPR001628 |
ENSP00000356959 |
T306714_2.00 |
GigaDB (2015-Oct-22) |
| NCBI Gene Info:This gene encodes a member of the nuclear receptor superfamily, and is a key regulator of xenobiotic and endobiotic metabolism. The protein binds to DNA as a monomer or a heterodimer with the retinoid X receptor and regulates the transcription of target genes involved in drug metabolism and bilirubin clearance, such as cytochrome P450 family members. Unlike most nuclear receptors, this transcriptional regulator is constitutively active in the absence of ligand but is regulated by both agonists and inverse agonists. Ligand binding results in translocation of this protein to the nucleus, where it activates or represses target gene transcription. These ligands include bilirubin, a variety of foreign compounds, steroid hormones, and prescription drugs. In addition to drug metabolism, the CAR protein is also reported to regulate genes involved in glucose metabolism, lipid metabolism, cell proliferation, and circadian clock regulation. Multiple transcript variants encoding different isoforms have been found for this gene. [provided by RefSeq, Jul 2020] |
Directly determined binding motifs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
| No direct experiments |
|
|
|
|
|
Motifs from related TFs |
|---|
| Name/Motif ID |
Species |
Forward |
Reverse |
Type/Study/Study ID |
SR
Score |
DBD
Identity |
NR1I3
M03924_2.00 |
Homo sapiens |
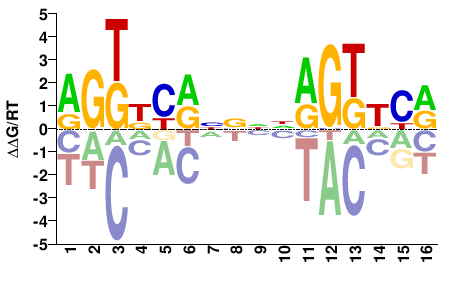
RGKDYRNNNNRGKKYR |
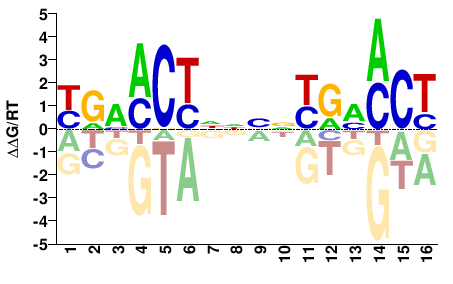

YRMMCYNNNNYRHMCY |
SELEX
Nitta et al.(2015)
NR1I3_1
|
0.992
|
0.986
|
NR1I3
M05619_2.00 |
Homo sapiens |
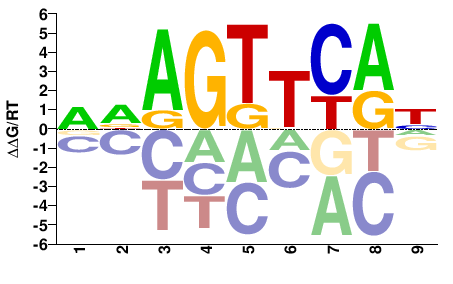
WDAGTTCAH |

DTGAACTHW |
SELEX
Yin et al.(2017)
NR1I3_FL_HT-SELEX
|
0.992
|
0.986
|
NR1I3
M09281_2.00 |
Homo sapiens |
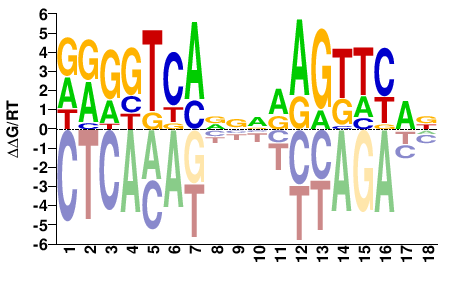
DRGGTCANNNRAGTTYAN |
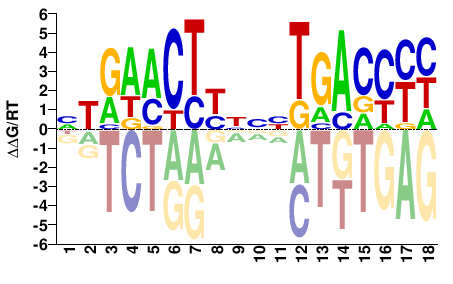
NTRAACTYNNNTGACCYH |
Misc
Kulakovskiy et al.(2013)
NR1I3_HUMAN.H11MO.0.C
|
0.992
|
0.986
|
NR1I3
M05620_2.00 |
Homo sapiens |
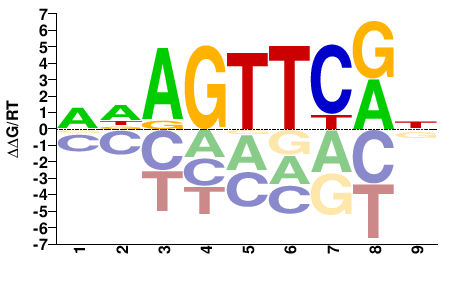
WDAGTTCRN |

NYGAACTHW |
SELEX
Yin et al.(2017)
NR1I3_FL_Methyl-HT-SELEX
|
0.992
|
0.986
|
Nr1i3
M09302_2.00 |
Mus musculus |
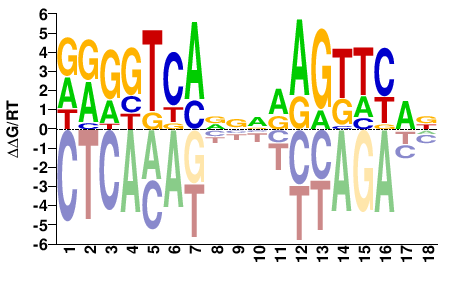
DRGGTCANNNRAGTTYAN |
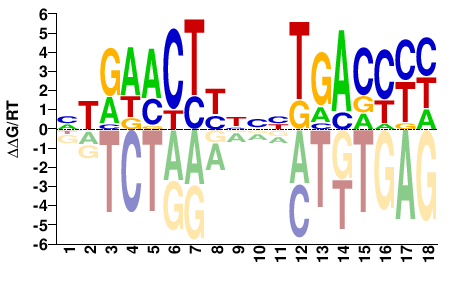
NTRAACTYNNNTGACCYH |
Misc
Kulakovskiy et al.(2013)
NR1I3_MOUSE.H11MO.0.C
|
0.985
|
0.886
|
ENSPMAG00000000180
M02403_2.00 |
Petromyzon marinus |

RGKWCRN |
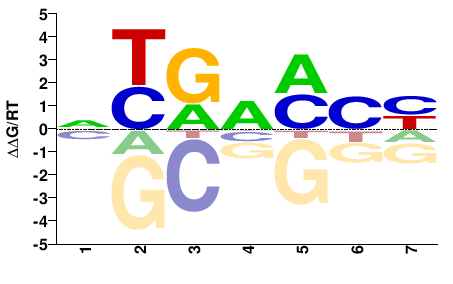
NYGWMCY |
PBM
Weirauch et al.(2014)
pTH5509
|
0.826
|
0.643
|
VDR
M03363_2.00 |
Homo sapiens |
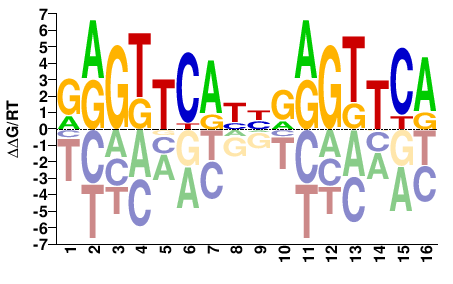
GRGTTCAYHGRGTTCA |
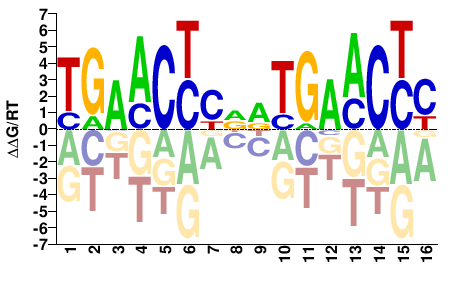
TGAACYCDRTGAACYC |
SELEX
Jolma et al.(2013)
VDR_1
|
0.816
|
0.629
|
Vdr
M03429_2.00 |
Mus musculus |
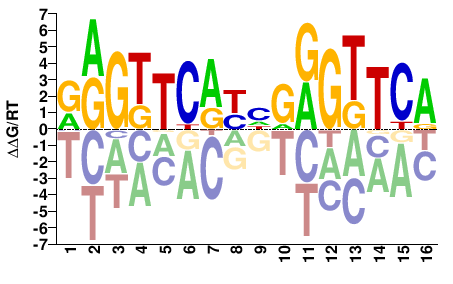
RRGTTCAYHGRGTTCA |

TGAACYCDRTGAACYY |
SELEX
Jolma et al.(2013)
Vdr_1
|
0.816
|
0.629
|
VDR
M05877_2.00 |
Homo sapiens |
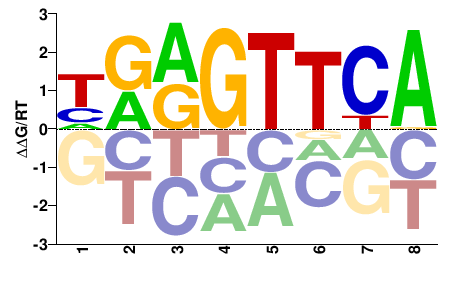

HRRGTTCA |

TGAACYYD |
SMiLE-seq
Isakova et al.(2017)
VDR
|
0.816
|
0.629
|
VDR
M09269_2.00 |
Homo sapiens |
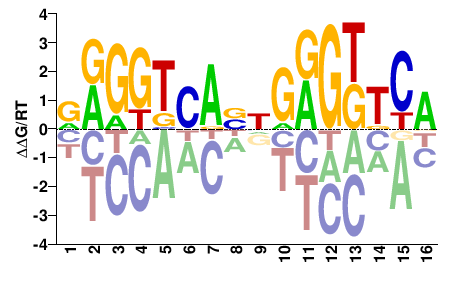
NRGGKCANNGRGKKCR |

YGMMCYCNNTGMCCYN |
Misc
Kulakovskiy et al.(2013)
VDR_HUMAN.H11MO.0.A
|
0.816
|
0.629
|
Vdr
M09315_2.00 |
Mus musculus |
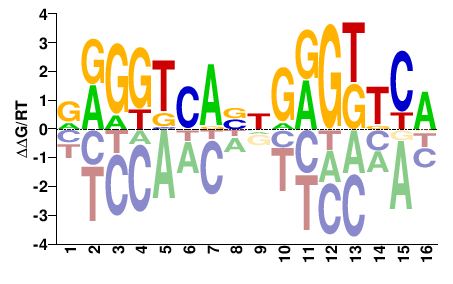
NRGGKCANNGRGKKCR |

YGMMCYCNNTGMCCYN |
Misc
Kulakovskiy et al.(2013)
VDR_MOUSE.H11MO.0.A
|
0.816
|
0.629
|
VDR
M09606_2.00 |
Homo sapiens |
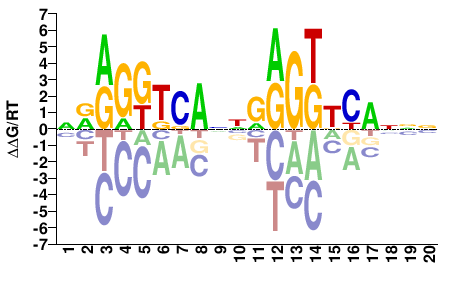
NRRGGTCANNRRGKKCANNN |
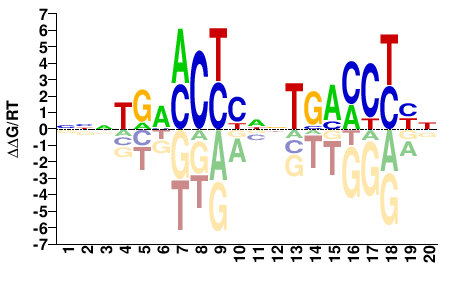
NNNTGMMCYYNNTGACCYYN |
Misc
Heinz et al.(2010)
GM10855-VDR+vitD_GSE22484
|
0.816
|
0.629
|
VDR
M11116_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$VDR_Q3
|
0.816
|
0.629
|
VDR
M11117_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$VDR_Q6_01
|
0.816
|
0.629
|
VDR
M11118_2.00 |
Homo sapiens |
Transfac license required
|
Transfac license required
|
Transfac
Matys et al.(2006)
V$VDR_Q6
|
0.816
|
0.629
|
NR1I2
M03925_2.00 |
Homo sapiens |
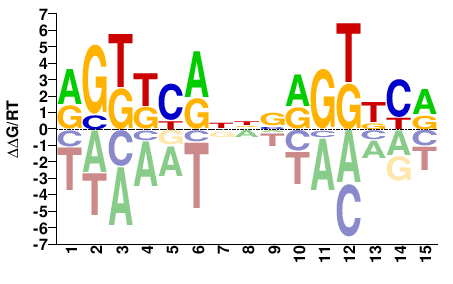
RGKKCRNNVRGKKCR |
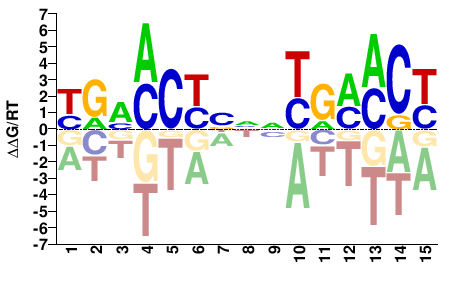
YGMMCYBNNYGMMCY |
SELEX
Nitta et al.(2015)
NR1I2_1
|
0.751
|
0.586
|
NR1I2
M05625_2.00 |
Homo sapiens |
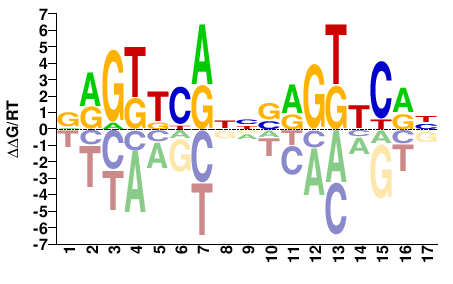
VRGTTCRNNSRGKTCRH |
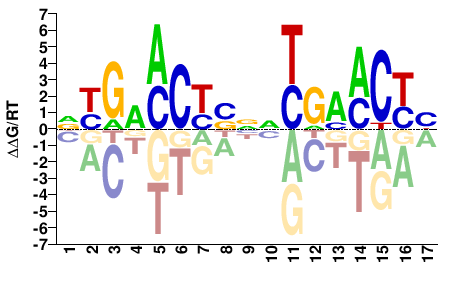

DYGAMCYSNNYGAACYB |
SELEX
Yin et al.(2017)
NR1I2_eDBD_HT-SELEX
|
0.751
|
0.586
|
NR1I2
M09283_2.00 |
Homo sapiens |
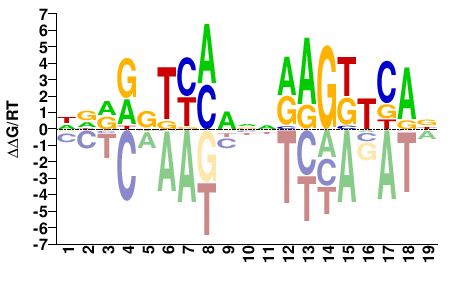

DDRRBTYMDNNRAGKTCAN |
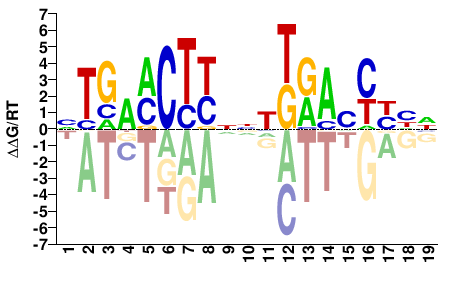
NTGAMCTYNNHKRAVYYHH |
Misc
Kulakovskiy et al.(2013)
NR1I2_HUMAN.H11MO.0.C
|
0.751
|
0.586
|
NR1I2
M05626_2.00 |
Homo sapiens |
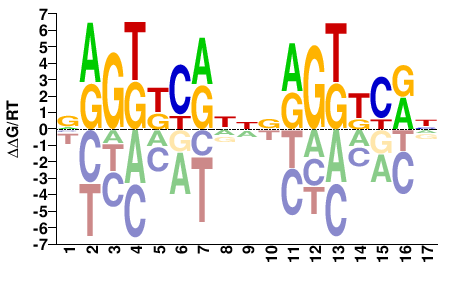
VRGKKCRNNVRGKKCRN |
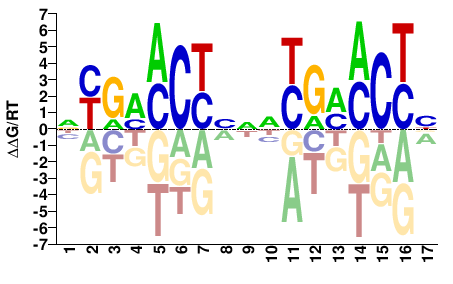
NYGMMCYBNNYGMMCYB |
SELEX
Yin et al.(2017)
NR1I2_eDBD_Methyl-HT-SELEX
|
0.751
|
0.586
|
| For this family, TFs with SR scores > 0.745 will likely have a similar motif |